Chapter: Genetics and Molecular Biology: Protein Synthesis
Verifying the Signal Peptide Model - Protein Synthesis
Verifying the Signal Peptide Model
Often it is easier to make observations on
eukaryotic cells and then to prove the resulting ideas with experiments done on
bacteria. One direct demonstration of concomitant translation and excretion
through a membrane was made in bacteria. Chains of a periplasmic protein in the
process of being synthesized were labeled by a chemical that was excluded from
the cytoplasm by the inner membrane. The experiment was done by adding the
labeling chemical to spheroplasts. These are cells lacking their outer membrane
and peptidyl-glycan layer. Shortly after adding the reactive labeling compound,
membrane-bound ribo-somes were isolated and their nascent polypeptide chains
were exam-ined and were found to be labeled. Proteins that are normally found
in the cytoplasm were not similarly labeled.
Does the N-terminal sequence on a protein signal
that the remainder of the protein is to be transported into or through a
membrane? In principle, this could be tested by tricking a cell into
synthesizing a new protein in which the N-terminal sequence from an excreted
protein has been fused to a protein normally found in the cytoplasm. If the
hybrid protein is excreted, then the new N-terminal sequence must be signaling
export.
Fortunately, E.
coli has been sufficiently well
studied that a number of candidates exist whose N-terminal sequences might be
used in such a project. The malF gene
product is a protein involved in the uptake of maltose into cells. It is
located in the periplasmic space. This should be an excellent source of an
“excretion-coding” N-terminal sequence. The ideal situation to test the
excretion hypothesis would be to fuse the N-terminal sequence of malF to an easily assayed cytoplasmic
protein. One very good candidate for this fusion is β-galactosidase, a protein for which the genetics have also been fully
developed.
Remarkably, the fusion of the N-terminal portion of
the malF gene to β-galactosidase was performed in vivo without using recombinant DNA techniques (Fig. 7.21).
Through clever genetic manipulations, Silhavy and Beckwith moved the β-galactosidase gene, lacZ, near malF and then
generated a deletion that fused the N-terminal portion of the malF gene toβ-galactosidase. A postulate of the signal peptide modelcould therefore
be tested. Indeed, a sizable fraction of the β-galactosi-dase from some of the fusion strains was not located in the
cytoplasm
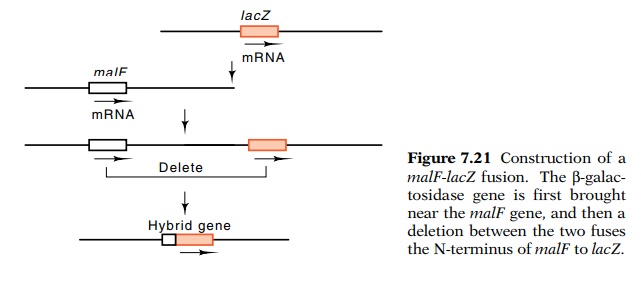
Figure
7.21 Construction of amalF-lacZ fusion. Theβ-galac-tosidase gene is first brought near the malF gene, and then a deletion between
the two fuses the N-terminus of malF
to lacZ.
How do you show that a protein is located in the
inner membrane of a cell? One method is to disrupt cells, to collect membrane
fragments, and then to separate the inner and outer membrane fragments and
assay each for the protein. Inner membrane is less dense than outer mem-brane,
and therefore the two can be separated according to density by isopycnic
centrifugation in a tube containing a gradient of sucrose concentration (Fig.
7.22). The two membrane fractions sediment down the tube into higher and higher
sucrose concentrations until they reach positions where their densities equal
the density of the sucrose. There they come to rest. By assay of the fractions
collected from such a sucrose gradient, the β-galactosidase from a malF-lacZ
fusion could be
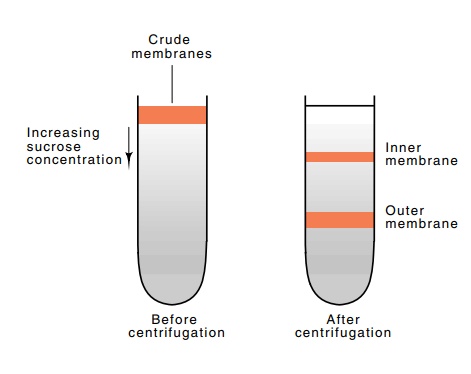
Figure
7.22 Isopycnic sepa-ration of membrane
fractions by centrifugation on a pre-formed sucrose gradient.
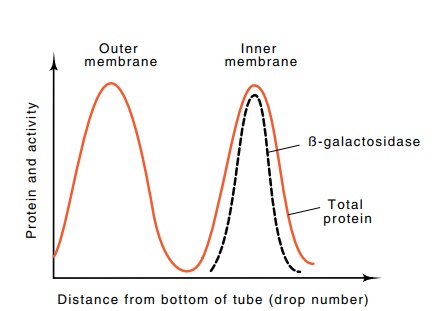
Figure
7.23 Results of isopycnic centrifugation
in which outer membranecame to rest near the bottom of the centrifuge tube and
inner membrane came to rest closer to the top. The β-galactosidase was associated with the inner membrane fraction.
Is the N-terminal sequence from malF actually necessary for export of
the β-galactosidase into or through the membrane? Some
of the fusions of malF to lacZ that were generated in the study
yielded proteins that remained in the cytoplasm. Genetic studies showed that
the process of producing the fusions in these strains had removed most of the
signal peptide of the malF gene.
Other studies have shown that mutations which abolish export of fusion proteins
alter the signal peptide. These experiments show that the signal peptide plays
an important role in export. It is clear, however, that the signal peptide is
not the whole story. The malF-lacZ
fusion protein was stuck to the inner membrane. It was not located in the
periplasmic space, the final destination of the native malF protein. Therefore something in the structure of theβ-galactosi-dase protein prevents the hybrid from
reaching the periplasmic space. Hence, reaching the periplasmic space requires
the leader sequence as well as a compatible structure in the remainder of the
protein.
Related Topics