Chapter: Genetics and Molecular Biology: Protein Synthesis
Base Pairing between Ribosomal RNA and Messenger
Base Pairing between Ribosomal RNA and Messenger
Ribosomes must recognize the start codons AUG or
GUG on the mes-senger RNA to initiate protein synthesis. Characterization of
the pro-teins that are synthesized when the lac
operon or other operons are induced show that normally only one AUG or GUG of a
gene is utilized to initiate protein synthesis. Many of the internal AUG or GUG
codons are not used to initiate protein synthesis. This means that something in
addition to the initiating codon itself must signal the point at which
translation begins.
Studies of bacterial translation show that the
first step in initiation is the binding of messenger to the smaller of the two
ribosomal subunits, the 30S subunit. The absence of a strictly conserved
sequence preceding start codons suggested that whatever first bound the
messenger to the 30S subunit might be an RNA-RNA interaction between mRNA and
ribosomal RNA. The originators of this idea, Shine and Dalgarno, were so
confident of their proposal that they proceeded to sequence the 3’ end of the
16S rRNA which is found in the smaller ribosomal subunit. They found that the
rRNA sequence provided strong support for their idea. The sequence on mRNA
which binds to the 16S ribosomal RNA is called the Shine-Dalgarno sequence or
the ribosome binding site.
Bacterial messengers contain a ribosome-binding
sequence slightly ahead of an initiating AUG. This sequence base pairs well
with a region
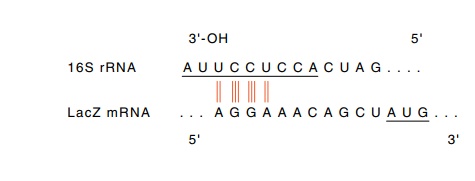
near the 3’ end of the 16S ribosomal RNA. This
upstream region has been examined in more than hundreds of messenger start
sequences. Typically it is three or four bases and is centered about ten
nucleotides ahead of the start codon. Despite the data to be described below,
the ribosome-binding sequence lying ahead of the AUG initiation codon is not
the whole story. Undoubtedly, secondary structure in the mRNA also can alter
translation efficiency. Occasionally, a sequence upstream or downstream from
the Shine-Dalgarno sequence pairs with it and blocks translation initiation.
Examination of the sequence in the vicinity of the AUG and ribosome-binding
sequence has shown the existence of preferences for some nucleotides. Were no
other factors involved, the identity of all these other nucleotides would be
random.
Related Topics