Chapter: Genetics and Molecular Biology: Protein Synthesis
The Signal Recognition Particle and Translocation - Protein Synthesis
The Signal Recognition Particle and
Translocation
In some types of eukaryotic cells, most of the
proteins translocated into or across membranes utilize the signal recognition
particles. These are small ribonucleoprotein particles containing a 300
nucleotide RNA molecule and five proteins, ranging in size from 14 KDa to 72
KDa. As the elongating signal peptide protrudes from the ribosome, the signal
recognition particles bind and arrest translation. After binding of the signal
recognition particle to the endoplasmic reticulum, translation resumes and the
protein is translocated across the membrane during its synthesis.
The generality of this process is not known, and some eukaryotic cells seem not to arrest translation.
Controversy has waxed hot on the matter of whether bacteria also utilize the
same sort of pathway. They contain a small ribonucleoprotein particle
containing an RNA that sediments at 4.5S. This possesses significant homology
to the eukaryotic signal recognition particle, but its role in protein
secretion is not yet clear.
Can the roles of the signal recognition particle
components be iden-tified? Genetics frequently can be utilized in simpler
organisms to obtain mutants unable to perform certain reactions. The complexity
of eukaryotic systems blocks the use of mutants for functional dissection of
the signal recognition particle. Instead, a biochemical approach had to be
used.
There are two requirements for a biochemical
dissection. First, it must be possible to assay for each of the steps in the
process. This is possible. Binding of signal recognition particles can easily
be measured because after binding they cosediment with the translating
ribosomes. By using homogenous messenger, translation arrest can be seen by the
failure of the proteins to be completed and by the accumulation of short
incomplete proteins. Translocation into membranes can be quantitated by adding
membrane vesicles to a translation mixture. Translocation of the protein into
the vesicles leaves them resistant to protease digestion.
The second requirement is the ability to take the
signal recognition particle apart and then to reassemble it. When the particle
had been dissociated and the various protein components were separated, they
were individually inactivated by reaction with N-ethylmaleimide. Par-ticles
were reassembled containing one reacted component and the remainder unreacted.
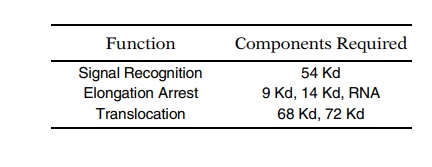
By this means, the proteins responsible for signal recognition, translation
arrest, and translocation were identified. In
Related Topics