Chapter: Biotechnology Applying the Genetic Revolution: Genomics and Gene Expression
Survey of the Human Genome
SURVEY
OF THE HUMAN GENOME
The sequence of the human genome
is 2.9 × 109 base pairs
(2.9 Gbp or gigabase pairs) in length. If the sequence were typed onto paper,
at about 3000 letters per page, it would fill 1 million pages of text. This
extraordinary amount of information is encoded by the sequence of just four
bases, cytosine, adenosine, guanine, and thymine. Most people expected the
human genome sequence to reveal the actual number of genes found in human
beings. In reality the massive amount of sequence needs sophisticated
interpretation in order to determine how many genes it contains. The best
estimates so far predict only 25,000 genes, but the number may be more or less.
Of the identified genes, we only know the function of around 50%. More than 40%
of the predicted human proteins are similar in structure to proteins in
organisms such as fruit flies or worms.
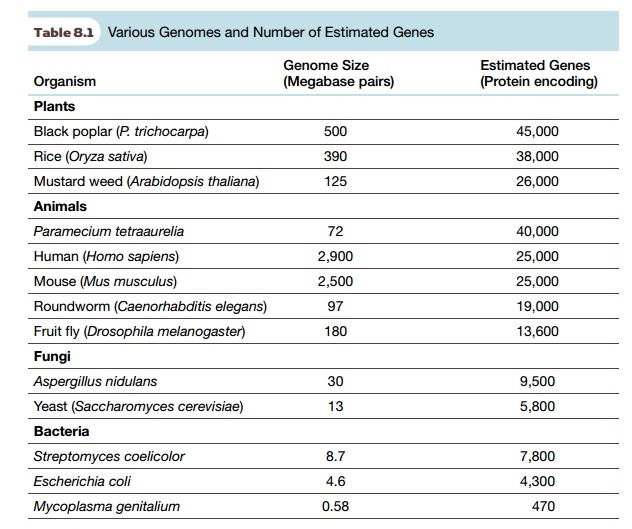
The genome sizes and estimated gene numbers are given for several organisms in Table 8.1. Note that plants presently hold the record for gene numbers, although they have less total DNA than higher animals. Among the animals, the ciliate protozoan Paramecium has less DNA but more genes than any multicellular animal sequenced so far. The largest bacterial genomes have more genes than the smaller eukaryotic genomes. An example is Streptomyces, famous as the source of many antibiotics. The smallest bacterial genomes, such as Mycoplasma, have fewer than 500 genes, although because these bacteria are parasitic they rely on the eukaryotes they infect for many metabolites.
In eukaryotic DNA, genes
encompass thousands or even millions of base pairs, most of which are actually
introns that are spliced out of the mRNA transcript. For example, the gene for
dystrophin (defective in Duchenne’s muscular dystrophy) is 2.4 million base
pairs long, and some of its introns are 100,000 base pairs or more in length.
In contrast, the coding sequence, consisting of multiple exons, is about 3000
base pairs. In such situations, it is not always easy to find coding sequences
among all the noncoding DNA. On the one hand, this may result in genes (or
individual exons) being completely missed. On the other hand, widely separated
exons that are in reality parts of a single coding sequence may be interpreted
as separate genes.
Another confounding factor in determining the number of genes is the presence of pseudogenes. These are duplicated copies of real genes that are defective and no longer expressed. They may be found next to the original, or they may be far away, on different chromosomes. Determining whether or not a “gene” is a pseudogene or genuine may be difficult with sequence data alone. Often the expression of a particular region of DNA must be confirmed by finding corresponding mRNA transcripts. DNA microarrays are a popular approach to confirming whether or not a gene is expressed (see later discussion).
The number of genes also
hinges on how we define a gene. In addition to the 25,000 protein-encoding
genes, there are a thousand or more genes that encode nontranslated RNA. The
ribosomal RNA and transfer RNA genes are the most familiar of these. However, a
variety of other small RNA molecules are involved in splicing of mRNA and in
the regulation of gene expression. Other sequences of DNA may not even be
transcribed, yet are nonetheless important. Should these also be regarded as
genes?
Related Topics