Chapter: Biotechnology Applying the Genetic Revolution: Genomics and Gene Expression
Sequencing Entire Genomes
SEQUENCING
ENTIRE GENOMES
Sequencing the entire genome from one organism can be accomplished in different ways. Chromosome walking allows the researcher to identify and sequence one clone and then, using those data, to find overlapping clones (Fig. 8.9). After those are identified and sequenced, more overlapping clones are identified. The process goes in order either up or down the chromosome, compiling the sequence piece by piece. Usually the first clone is located relative to a particular marker, such as an STS or RFLP. Chromosome walking is often used to characterize genes responsible for a particular disease. Analysis of DNA from people with the disease may have revealed a particular RFLP that is always present in those with the disease, but absent in unaffected people. This RFLP can be identified in a library clone. Then chromosome walking both upstream and downstream of the RFLP will, it is hoped, provide the whole gene sequence.

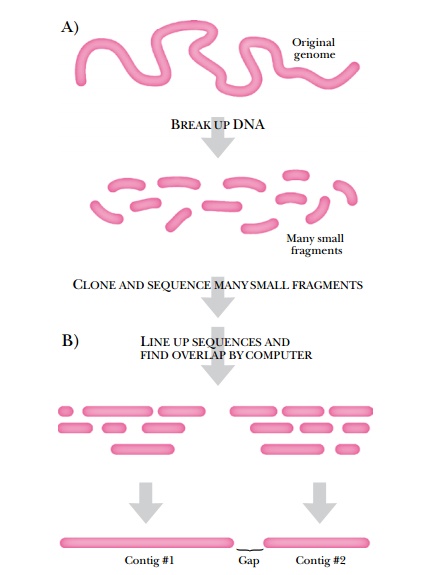
Although chromosome walking is a powerful tool to identify genes, its use for sequencing an entire genome is too arduous. Instead, shotgun sequencing is used to assemble sequence data from an entire genome (Fig. 8.10). Here genomic libraries are constructed and random clones are sequenced. A computer compiles the sequence information, identifying the overlapping regions between clones, and ordering the clones into a complete sequence. This procedure is repeated to eliminate as many gaps as possible. This method was used to sequence Haemophilus influenzae, the first cellular genome to be sequenced. This genome has 1.8 Mb and took less than 3 months to complete using shotgun sequencing.

Related Topics