Chapter: Biotechnology Applying the Genetic Revolution: Genomics and Gene Expression
Genetic Mapping Techniques
GENETIC
MAPPING TECHNIQUES
Genomes are sequenced by
making libraries of genomic DNA segments and then sequencing each of the
segments. These stretches must then be compiled into the final sequence. To
structure the sequence data into a draft genome, the Human Genome Project
started by compiling a working genome map. Genome
maps provide various landmarks for use when putting together sequence data.
There are two different categories for genome maps, genetic maps and physical maps. Genetic maps are based
on the relative order of genetic markers,
but the actual distance between the markers is hard to determine. Physical
maps are more precise and give the distance between markers in base pairs.
Traditional genetic maps are
based on the recombination frequency between genes.
In eukaryotic cells,
recombination occurs between homologous pairs of chromosomes during meiosis. If
two genes are close together on the same chromosome, recombination between them
will be rare. If the two genes are located far apart on the same chromosome,
recombination is relatively frequent. Early genetic maps were based on
measuring recombination frequencies between genes.
Genetic maps are based on
landmarks called genetic markers.
Many different types of markers can be used. The order of these markers is
determined by how often the two markers are found in offspring. The most useful
markers are genes, but often in large genomes, genuine genes are too few and
far between to give a good map. Genes that encode specific traits are wonderful
markers in organisms such as Drosophila
because mating can be controlled and directed. After mating many different
flies, the number of flies with both markers can be determined. The more often
the markers appear together in the offspring, the closer these are in the
genome. In humans, deliberate mating experiments are unethical. Moreover, the
human genome contains only a few percent of coding DNA; thus, using real genes
does not produce enough points on the map. A sparse map makes it difficult to
order the sequences obtained in the genome sequencing project. Therefore, other
markers, including physical markers, are also used on genomic maps.
An example of a physical
marker is the RFLP, or restriction fragment length polymorphism. RFLPs are
commonly used because of the ease of identification. For small genomes such as
yeast, monitoring the frequency of recombination between two RFLP markers is
easy. Diploid yeast cells undergo meiosis and form four haploid cells called a
tetrad. Each of these haploid cells can be isolated, grown into many identical
clones, and examined individually. Thus each RFLP marker can be followed easily
from one generation to the next. In humans, following such markers is more
challenging, but studies on groups of closely related people, such as large
families or small cultures like the Amish, have allowed some RFLPs to be
followed in this manner (Fig. 8.1).
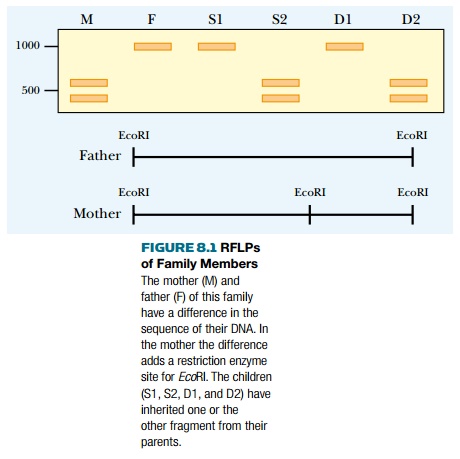
Another marker used for
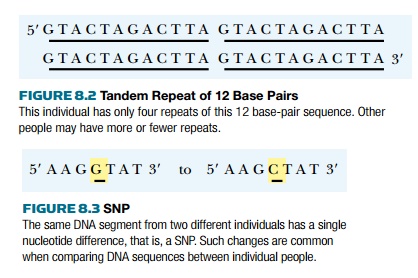
making genetic maps is the VNTR, or variable number tandem repeat (Fig. 8.2).
These sequence anomalies occur naturally in the genome and consist of tandem
repeats of 9 to 80 base pairs in length. The number of repeats differs from one
person to the next; therefore, these can be used as specific markers on a
genetic map. They can also be used to identify individuals in forensic medicine
or paternity testing. Some repeats are found in many different locations
throughout the genome and cannot be used for making genetic maps, but other
repeat sequences are found only in one unique location.

A third type of marker is the
microsatellite polymorphism, which is also a tandem repeat. However, unlike VNTRs, microsatellite
polymorphisms are repeats of 2 to 5 base pairs in length, and usually consist
of cytosine and adenosine.
A fourth type of genetic
marker used in mapping is the SNP (pronounced “snip”), or single nucleotide
polymorphism (Fig. 8.3). SNPs are individual substitutions of a single
nucleotide that do not affect the length of the DNA sequence. These changes can
be found within genes, in regulatory regions, or in noncoding DNA. When found
within the coding regions of genes SNPs may alter the amino acid sequence of
the protein. This in turn may affect protein function. If a SNP correlates with
a genetic disease, identifying that SNP may diagnose the disease before
symptoms appear. When a SNP falls within a restriction enzyme site, it
coincides with an RFLP.
Related Topics