Chapter: Biochemistry: Photosynthesis
Chloroplast Genes
Chloroplast
Genes
Chloroplasts,
like mitochondria, have their own DNA. Scientists speculate that these
organelles were originally cyanobacteria that were engulfed by a eukaryotic
cell through endosymbiosis. Some interesting, even elaborate, interactions have
evolved from the interplay of both nuclear and chloroplast genes. There are
about 3000 chloroplast proteins, but 95% of them are encoded by nuclear genes.
One of
the interesting interactions involves rubisco, the prin-cipal enzyme of CO2
fixation. The large subunit of this enzyme is coded by a chloroplast gene, and
the small subunit by a nuclear gene. Mechanisms not yet understood must
coordinate the two syntheses to ensure equimolar production of both subunits.
The
nuclear gene is translated in the cytoplasm, and the protein is then
transported to the chloroplast, protected by a chaperonin, via targeting
mechanisms. Special targeting sequences are used to direct various nuclear
products to the appropriate chloroplast location, using reactions that require
ATP hydrolysis. The chaperonin aids in formation of the final, active complex.
The
chloroplast encodes its own RNA polymerase, ribo-somal and transfer RNAs, and
about one-third of the ribosomal proteins.
The DNA
polymerase, aminoacyl synthetases, and the rest of the ribosomal proteins come
from the nuclear genes. Different nuclear genes may be used for chloroplasts in
different, special-ized tissues of the plant. In different classes of plants,
differ- ent genes are coded by the nucleus, even though all the same proteins
constitute the chloroplast. The sequences of many of these enzymes encoded in
the nucleus resemble those found in bacteria more than the sequences of
proteins encoded by other nuclear genes. The chloroplast rRNAs likewise
resemble bacte-rial rRNAs. The four subunits of the chloroplast-coded RNA
polymerase are homologous to the four subunits of bacterial RNA polymerase.
Furthermore, the chloroplast mRNA uses a Shine-Dalgarno sequence to bind to the
ribosome, and it does not have a cap or a poly-A tail. These observations are
definitely consistent with the idea that the chloroplast (and mitochondri-on)
originated from bacteria-like organisms, symbiotically taken into early cells,
with some subsequent transfer of the organelle genes to the nucleus.
The
separation of genetic material between the nucleus and the chloroplast requires
a coordination of the transcription of genes in two different locations. How
plant cells are able to accomplish this is an active research area. It is now
known that there are communications in both directions between the two
organelles. A signal proceeding from the nucleus to the chloro-plast is
considered the more ÒnormalÓ pathway and is referred to as anterograde signaling. When the signal goes from another organelle
to the nucleus, it is called retrograde
signaling. Recently, three different retrograde signaling processes were
discovered in algae and higher plants.
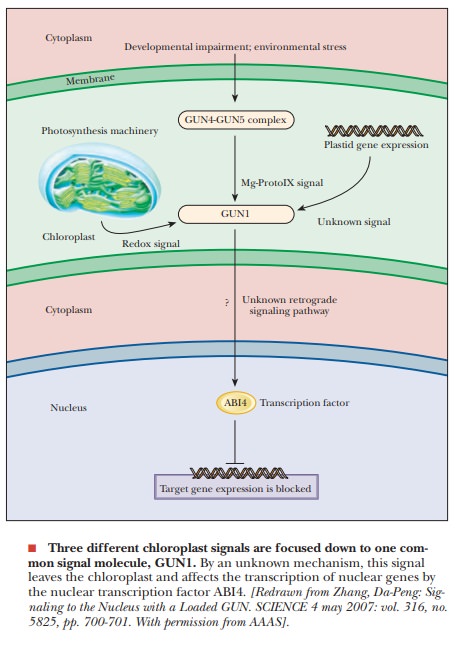
The
most-studied is signaling by Mg-protoporphyrin IX, a tetrapyrrole generated
during chlorophyll biosynthesis. Also studied are signals coming from the
expression of chloroplast genes and those from the photosynthetic electron
transport chain (PET). Stress signals and developmental impairment from the
cytosol increase the levels of another set of related proteins, GUN4 and GUN5.
These three retrograde systems have been shown to act through a GUN1 (genomes
uncoupled 1), which is a member of a large family of enigmatic transcription
factors in higher plants. It has been shown that all three signal pathways
affect the level of GUN1 in the chloroplast (see the figure). It is currently
unknown how the GUN1 signal makes its way to the nucleus. However, in the
nucleus, the effect is to increase a transcription factor called ABI4 (abscisic
acid insensitive 4), which then inhibits transcription of targeted nuclear
genes involved in chloroplast metabolism. This common pathway focused to the
GUN1 protein allows the smooth coordination of the nuclear part of chloroplast
metabolism in response to many environmental signals.
Related Topics