Chapter: Biotechnology Applying the Genetic Revolution: RNA-Based Technologies
RNAi for Studying Mammalian Genes
RNAi
FOR STUDYING MAMMALIAN GENES
An important application for
RNAi is testing the individual roles of human proteins, which can reveal new
targets for curing diseases. Until recently, using RNAi to assess the function
of mammalian proteins was not possible. The application of dsRNA to cultured
mammalian cells or whole mice induces a potent antiviral response. Interferon
is produced, which triggered the cells to degrade all RNA transcripts and shut
down protein synthesis. Thus, the methods used in C. elegans and Drosophila
killed mammalian cells, and no alternatives were known until recently.
Recognizing that short interfering
RNA (siRNA) pieces mediated RNAi was the key to its application in mammalian
systems. Instead of using long dsRNA as in C. elegans and Drosophila, exposing
mammalian cells to dsRNA shorter than 30 nucleotides activated the mammalian
counterparts to Dicer and RISC. This in turn abolished expression of the target
mRNA. Such short dsRNA segments thus act as endogenously produced siRNA. As
described earlier, siRNAs made by Dicer are short double-stranded RNA about 21
to 23 base pairs in length. In addition, the siRNAs have a two-base 3′ overhang
that is more stable when it consists of two uracils. To study a particular
target mRNA in mammalian cells, chemically synthesized siRNAs with these
characteristics are designed to have the complementary sequence to the target.
Much like antisense oligonucleotides, these may have modifications to make them
more stable, such as methyl groups added to the 2′-OH of the ribose. The most important and the
most difficult aspect of in vitro
siRNA construction is determining an effective sequence. The sequence on the
target mRNA must be accessible to the siRNA, which may be challenging because
of RNA secondary structure. Many suitable siRNA are designed and synthesized,
and each is tested for activating RNAi. These siRNA are delivered to mammalian
cells much like antisense oligonucleotides, including transfection, liposomes,
and microinjection.
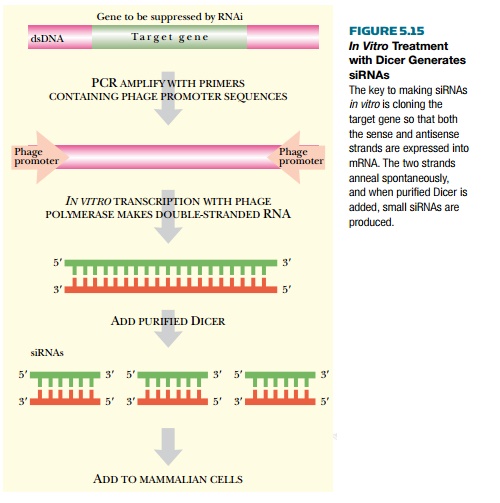
Rather than chemically
synthesizing siRNA, the target mRNA can be mixed with purified Dicer to cleave
the target mRNA into siRNA pieces (Fig. 5.15). Although the reaction occurs in
a tube, the purified Dicer can generate multiple siRNAs as it would in vivo. In order to create the target
mRNA, the chosen target gene is amplified with PCR using PCR primers that also
contain a promoter sequence for T7 RNA polymerase. The double-stranded DNA is
then converted into dsRNA by T7 RNA polymerase. The RNA pieces are allowed to
anneal spontaneously, and then they are mixed with purified Dicer, which
digests it into multiple different siRNAs. These are then transfected into
mammalian cells.
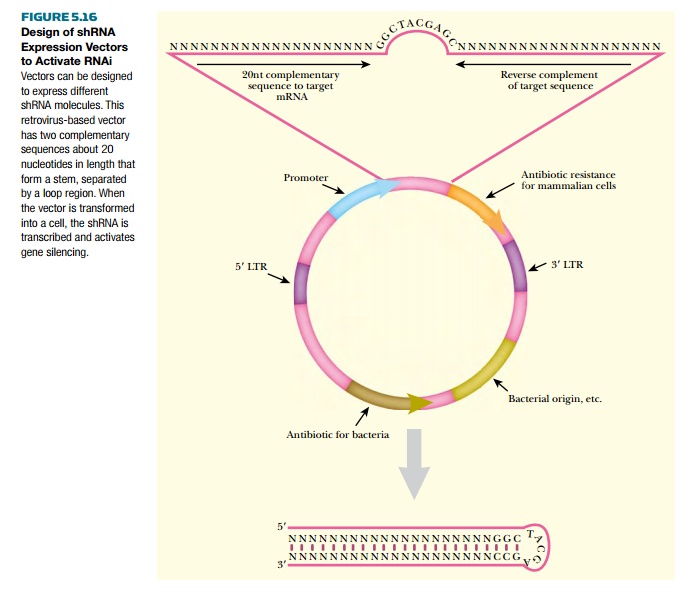
Additionally, mammalian cells can be induced to activate RNAi by expressing short hairpin RNAs (shRNAs) that mimic the structure of microRNAs. A gene for the shRNA is constructed in a vector. The shRNA can be transcribed as two complementary strands by two different promoters facing in opposite directions, or simply made as one transcript with complementary ends interrupted by a sequence that forms a loop (Fig. 5.16). In both constructs, the complementary sequences come together to form a double-stranded RNA region. The most common promoter for expressing shRNA in vivo is the promoter for mammalian RNA polymerase III. (Note: This enzyme normally transcribes small noncoding RNAs.) RNA polymerase III starts transcription at a specific sequence and stops transcription when it encounters four to five consecutive thymidines. In addition, RNA polymerase III does not activate the enzymes that cap and add a poly(A) tail onto the transcript. This polymerase is precisely what is necessary to create an shRNA that mimics one found in eukaryotic cells.
Rather than designing the
shRNA constructs from scratch, another strategy uses preexisting microRNA.
First, the stem portion is
replaced with sequences that match the target mRNA. The newly designed microRNA
will trigger RNAi for the target mRNA rather than its endogenous target.
Vectors that express shRNAs
have some advantages over adding siRNA. When siRNA is added to mammalian
cultures, the effect is temporary. When the siRNA is gone, the effect ends
(unless heterochromatin formation was triggered, as seen in some organisms).
When an shRNA vector is transfected into a mammalian cell, the effect is more
sustained. The vector continues to produce shRNA for a considerable time. In
addition, expression of the shRNA may be controlled using promoters that are
inducible or tissue-specific. In a clinical setting, the ability to deliver the
siRNA only to those tissues that need it is crucial, and using a vector system
can accomplish this.
Related Topics