Chapter: Biochemistry: The Three-Dimensional Structure of Proteins
Denaturation and Refolding
Denaturation and Refolding
The noncovalent interactions that maintain the three-dimensional structure of a protein are weak, and it is not surprising that they can be disrupted easily.
The unfolding of a protein (i.e., disruption of the tertiary structure) is called denaturation. Reduction of disulfide bonds leads to even moreextensive unraveling of the tertiary structure. Denaturation and reduction of disulfide bonds are frequently combined when complete disruption of the tertiary structure of proteins is desired. Under proper experimental conditions, the disrupted structure can then be completely recovered. This process of denaturation and refolding is a dramatic demonstration of the relationship between the primary structure of the protein and the forces that determine the tertiary structure. For many proteins, various other factors are needed for complete refolding, but the important point is that the primary structure determines the tertiary structure.
Proteins can be denatured in several ways. One is heat. An increase in tem-perature favors vibrations within the molecule, and the energy of these vibra-tions can become great enough to disrupt the tertiary structure. At either high or low extremes of pH, at least some of the charges on the protein are missing, and so the electrostatic interactions that would normally stabilize the native, active form of the protein are drastically reduced. This leads to denaturation.
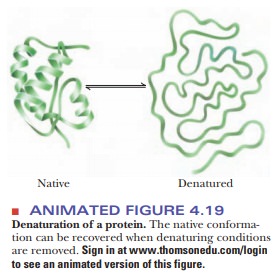
The binding of detergents, such as sodium dodecyl sulfate (SDS), also dena-tures proteins. Detergents tend to disrupt hydrophobic interactions. If a deter-gent is charged, it can also disrupt electrostatic interactions within the protein. Other reagents, such as urea andguanidine hydrochloride, form hydrogen bonds with the protein that are stronger than those within the protein itself. These two reagents can also disrupt hydrophobic interactions in much the same way as detergents (Figure 4.19).
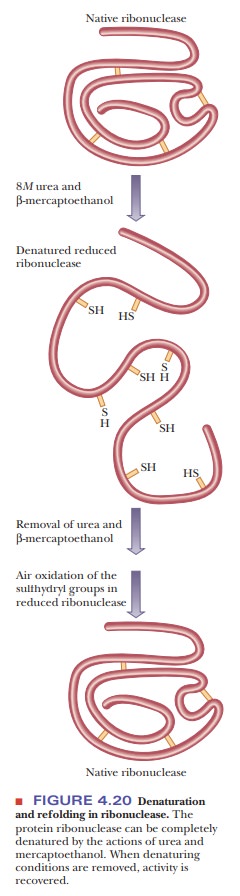
b-Mercaptoethanol (HS-CH2-CH2-OH) is frequently used to reduce disul-fide bridges to two sulfhydryl groups. Urea is usually added to the reaction mixture to facilitate unfolding of the protein and to increase the accessibility of the disulfides to the reducing agent. If experimental conditions are properly chosen, the native conformation of the protein can be recovered when both mercaptoethanol and urea are removed (Figure 4.20). Experiments of this type provide some of the strongest evidence that the amino acid sequence of the protein contains all the information required to produce the complete three-dimensional structure. Protein researchers are pursuing with some interest the conditions under which a protein can be denatured-including reduction of disulfides-and its native conformation later recovered.
Related Topics