Chapter: Biochemistry: Nucleic Acid Biotechnology
The Polymerase Chain Reaction
The Polymerase Chain Reaction
It is possible to increase the amount of a given DNA many times
over without cloning that DNA. The method that makes this amplification
possible is the polymerase chain
reaction (PCR). Any chosen DNA can be amplified, and it neednot be
separated from the rest of the DNA in a sample before the procedure is applied.
PCR copies both complementary strands of the desired DNA sequence. Scientists
had long wished for a cell-free, automated way of synthesizing DNA, but any
system that could be automated would need to function at high temperatures so
that the DNA strands could be separated physically without the need for the
many enzymes found in DNA replication, such as topoisomerases and helicases.
Unfortunately, such temperatures (around 90°C) would denature and inactivate
the DNA polymerases that would be making the DNA strands. What made the process
possible was the discovery of bacteria that live around deep-sea hydrothermal
vents, under extreme pressures, and at temperatures higher than 100°C. If
bacteria can live under those conditions, then their enzymes must be able to
function at those temperatures. The bacterium, Thermus aquaticus, from which a heat resistant polymerase is
extracted, is one ofthose that live in hot springs. The enzyme is called Taq polymerase. We saw that the
biotechnology industry eagerly searches for organisms that live under extreme
conditions, and here we have an example of why it does so.

At the start of the process, the two DNA strands are separated by heating, after which short oligonucleotide primers are added in large excess and, via cooling, are allowed to anneal to the DNA strands. These primers are complementary to the ends of the DNA chosen for amplification and serve the same purpose as the RNA primers in normal replication. Once the primers have annealed to the DNA, the temperature is raised again to optimize the activity of Taq polymerase, which begins synthesizing the new DNA from the 3' end of the primer. The two complementary strands grow in the 5' to 3' direction (Figure 13.23), and the Taq polymerase is allowed to work until the desired length of DNA has been synthesized.
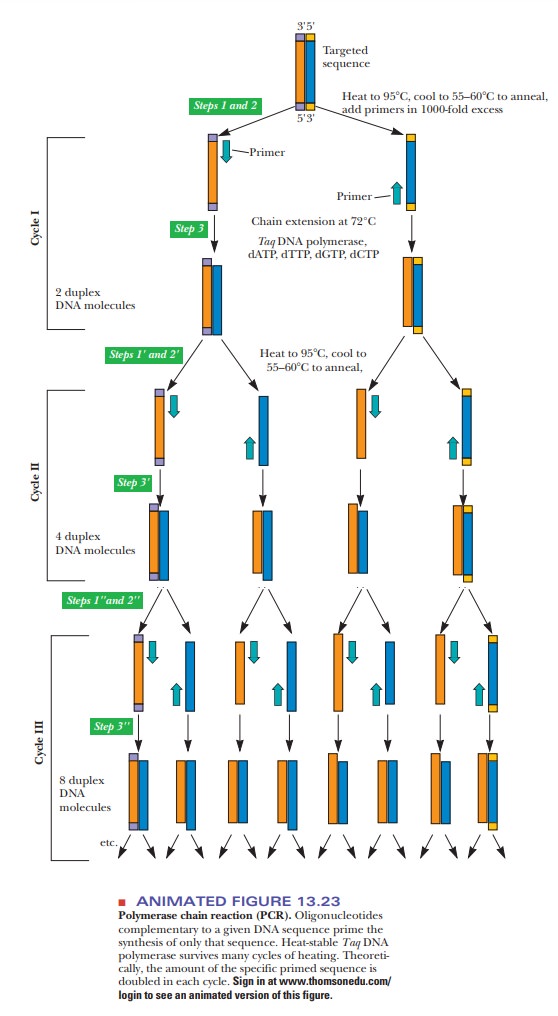
This first round doubles the amount of the desired DNA. The
process of unwinding the two strands, annealing the primers, and copying the
complementary strand is repeated, bringing about a second doubling of the
selected double-stranded DNA. It is not necessary to add more primer because it
is present in large excess. The whole process is automated. Control of the
temperature to which the strands are heated to separate them is crucial, as is
the temperature chosen for annealing the primers.
The amount of DNA continues to double in subsequent rounds of
ampli-fication. After about an hour, and 25 to 40 cycles of replication, one
obtains millions to hundreds of millions of copies of the desired DNA segment,
usually a few hundred to a few thousand base pairs long (Figure 13.23). Other
DNA sequences are not amplified and do not interfere with the reaction or
subse-quent use of the amplified DNA.
The most important part of the science behind PCR is the design of
the primers. They have to be sufficiently long to be specific for the target
sequence but not so long that they are too expensive. Usually, the primers are
18 to 30 bases long. They must also have optimal binding properties, such as an
amount of G and C that is sufficient to allow them to anneal before the entire
DNA renatures. In addition, the two primers should contain similar amounts of G
and C so that they have the same melting temperature. The sequence of the
primer must not lead to secondary structures within a primer or between the two
different primers; otherwise, the primers bind to themselves instead of to the
DNA being amplified. For example, if a primer had the sequence AAAAATTTTT, it
would form a hairpin loop with itself and would not be avail-able to bind the
DNA.
What are the advantages of PCR?
Amplification of the amounts of DNA in extremely small samples has
made it possible to obtain accurate analyses that were not possible earlier.
Forensic applications of the technique have resulted in positive
identifications of crime victims and suspects. Even minuscule amounts of
ancient DNA, such as those available from Egyptian mummies, can now be
researched after amplification. The following Biochemical Connections box
describes some forensic uses of DNA technology.
Summary
Polymerase chain reaction (PCR) is a sophisticated, automated
technique for amplifying DNA from very small amounts of sample.
The DNA to be amplified is mixed with specific primers, dATPs,
dCTP, dGTP, dTTP, and a heat-stable form of DNA polymerase.
The mixture undergoes 20 to 40 rounds of DNA polymerization via
cycling the temperatures so that the DNA strands separate, the primers anneal,
and the polymerase fills in the DNA. Each cycle doubles the target DNA.
The technique has revolutionized forensic science, as DNA can be
ampli-fied from just a few cells and then the DNA analyzed and identified.
Related Topics