Chapter: Biochemistry: Nucleic Acid Biotechnology
Fusion Proteins and Fast Purifications
Fusion Proteins
and Fast Purifications
Affinity chromatography was introduced as an example of a powerful
technique for protein purification. Molecular biologists have taken the idea
one step further and have incorporated affinity chromatography ligand-binding
sites directly into a protein to be expressed. A protein is created that
contains not only the amino acid sequence of the desired polypeptide but also
some extra amino acids at the N-terminus or C-terminus. These new proteins are
called fusion proteins. The figure
indicates how this might work. An expression vector that has a promoter for T7
polymerase, followed by a start sequence ATG, is used. A his-tag sequence that follows the ATG codes for six histidine
residues. Following the his-tag is a sequence that is specific for a
proteolytic enzyme called enterokinase.
Finally comes the MCS, where the gene of interest can be cloned. After the
desired
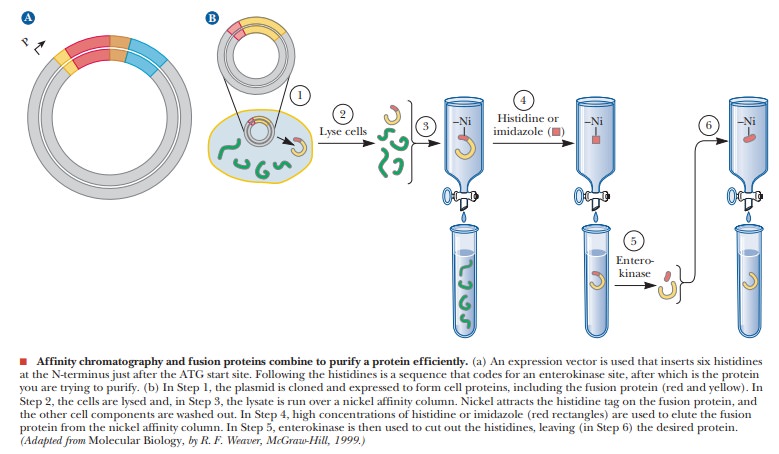
The fusion proteins that are translated have the initial
methionine, six histidines, the enterokinase-specific amino acid sequence, and
then the desired protein. This technique is used with a his-tagged fusion
protein. A nickel affinity column is set up, which is very specific for
histidine residues. The cells are lysed and passed over the column. All the
proteins pass through except the fusion protein, which binds tightly to the
nickel column. The fusion protein can be eluted with imidazole, a histidine
analog. Enterokinase is then added to cleave off the his-tag, leaving the
desired protein. Under ideal circumstances, this can be an almost perfect,
one-step purification of the protein.
Related Topics