Chapter: Basic Concept of Biotechnology : Tools and Techniques in Biotechnology
Microarray - Techniques of Biotechnology and Innovations
MICROARRAY
Microarray technology is used to monitor genome wide expression levels of genes in a known organism. In this technology, aglass slide is used on to which DNA molecules are fixed in an organized manner at definite locations called spot. It may contain thousands of spots and these spots may hold a few million copies of identical DNA molecules that exclusively belong to a gene (Fig. 11A). The DNA in a spot may either be genomic DNA or short stretch of oligonucleotide strands that correspond to a gene. The spots or features are synthesised by the process of photolithography or printed on to the glass slide by a robot.
Microarrays are used to measure gene expression in several ways. One of the several popular applications is to compare the expression of a set of genes from a cell, to be diagnosed, (condition A) to the same set of genes from a reference cell maintained under standard conditions (condition B). Figure 11B provides a general picture of the involved experimental steps. First, RNA is extracted from the cells and reverse transcribed into cDNA by using reverse transcriptase enzyme and nucleotides labelled with different fluorescent dyes. cDNA from cells developed in condition A may be labelled with a red dye and from cells developed in condition B with a green dye. After differentially labelling the samples, they are hybridized onto the same glass slide. Any cDNA sequence in the sample hybridizes to specific spots present on the glass slide with its complementary sequence. The amount of cDNA bound to a spot is found to be directly proportional to the initial no. of RNA molecules available for that gene in both the samples.
The spots in the hybridized microarray are made to excite by a laser and scanned at appropriate wavelengths to distinguish the red and green dyes. The amount of fluorescence emitted upon excitation corresponds to the amount of nucleic acid bound. If cDNA from condition A for a particular gene is in greater quantity than that from condition B, the spot is found to be red. If cDNA from condition B is in abundance, the spot would be green. If the gene is expressed equally in both conditions, yellow spot is found, and if the gene is not expressed ineither condition, black spot is found. Thus, the end of the experiment is an image of the microarray where each spot corresponding to a gene has an associated fluorescence value which represents the relative expression level for that gene.
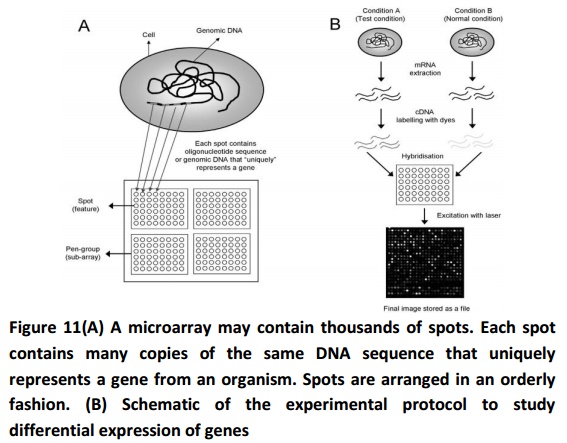
The organism is grown in two different conditions (a reference condition and a test condition). RNA is extracted from the two cells, and is labelled with different dyes (red and green) during the synthesis of cDNA by reverse transcriptase. This step is followed by hybridization of cDNA onto the microarray slide, where each cDNA molecule representing a gene will bind to the spot which contains its complementary DNA sequence. The microarray slide is then exposed to a laser for excitement at suitable wavelengths to detect the red and green dyes. The final image obtained is stored as a file for additional study.
Application of Microarrays
· Sensitive enough to detect single base differences, SNPs or mutations
· Useful for a wide range of applications viz. identifying contamination of food products with cells from other animals or plants; identifying strains of viruses; detecting mutations in cancer cells of a patient that may influence the disease’s response to treatment.
· Protein microarrays are developed for massive screening of interactions between proteins on the microarray, and other proteins, ligands or substrates.
Related Topics