Chapter: Basic Concept of Biotechnology : Tools and Techniques in Biotechnology
Fluorescence In Situ Hybridization (FISH) - Techniques of Biotechnology and Innovations
FLUORESCENCE IN SITU HYBRIDIZATION (FISH)
FISH is used to detect small deletions and duplications that are not visible for microscope analysis. It is used to detect the presence of no. of chromosomes of a certain type in each cell or to substantiate rearrangements that are visible in microscope analysis. FISH is used specifically to look at a specific area of one chromosome only. It uses A very small chemical is used which glows brightly after detecting a specific region on the chromosome. A special microscope is used to look at the chromosome to find out the no. of bright spots. In the case of deletion, only one bright spot is visible instead of two (one on each chromosome) whereas in the case of a duplication, three bright spots are found instead of just two.
FISH uses probes, small DNA strands, which have a fluorescent label attached to them. These probes are designed to be complementary to specific parts of a chromosome. When DNA is denatured on heating, the probes hybridise (Fig 7) to their complementary sequence in the DNA of the patient. The probe will not hybridise if a small deletion is found in the region complementary to the probe. More no. of probe hybridise if duplication is present.
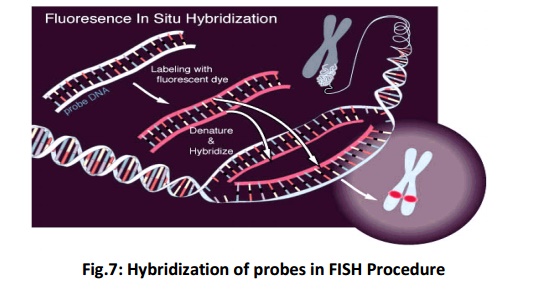
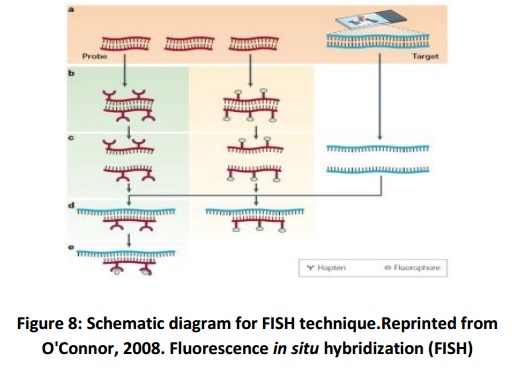
These are the following steps in the FISH procedure (Fig. 8):
i) Denaturing of the chromosomes
ii) Denaturing of the probe
iii) Hybridization
iv) Fluorescence staining
v) Examining the slides or storing in the dark
FISH testing for a deletion
Two types of probes are generally used; the first probe (generally green) is known as control probe which identifies both copies of the chromosome under test. It gets hybridised to a sequence that is not in the deletion region, so a signal is seen on each chromosome. The second probe (generally red) gets hybridised to the sequence that may be deleted. A deletion is generally present in only one of the chromosomes in a pair, therefore the probe bind to the undamagedchromosome, but is incapable to bind to the deleted chromosome which results in only one signal.
Types of probes
Three different types of FISH probes are used. Each of them has a different application:
Locus specific probes –These probes bind to a particular region of achromosome and is used when a small portion of a gene is isolated to determine on which chromosome the gene is located.
i) Centromeric repeat probes –Alphoid or centromeric repeat probesare the result of repetitive sequences present in the middle of each chromosome. These probes are used to determine whether an individual has the correct number of chromosomes. These probes are also used in combination with locus specific probes for determining missing genetic material from a particular chromosome in an individual.
ii) Whole chromosome probes –These are actually collections ofsmaller probes. Each of the smaller probes binds to a different sequence along the length of a given chromosome. They are particularly useful for examining chromosomal abnormalities viz. when a piece of one chromosome is found to be attached to the tail of another chromosome.
· Samples for FISH testing:
FISH is mostly performed on blood samples from both, adults and children. FISH is also used as a prenatal test for aneuploidy, extra copies of whole chromosomes, using placental samples from chorionic villus sampling (CVS) or amniotic fluid from an amniocentesis. It is also used as a prenatal test for deletions.
· Limitations of FISH:
FISH probes are available for the well characterized duplication and deletion syndromes only. It is difficult to see a duplication using FISH because the attachment of extra probes is not very easy to observe.
Applications of FISH in Clinical Practice
i) Pre-implantation Diagnostics
ii) Prenatal Diagnostics
iii) Tumor Diagnostics
iv) Postnatal Diagnostics
v) Centromere Mutations
Related Topics