Chapter: Genetics and Molecular Biology: Genetic Engineering and Recombinant DNA
Walking Along a Chromosome to Clone a Gene
Walking Along a Chromosome to Clone a Gene
Another method of cloning a desired gene is
walking. Suppose a randomly isolated clone has been shown to lie within
several hundred thousand bases of the DNA segment we wish to clone. Of course,
demonstrating such a fact is not trivial in itself. In the case of the fruit
fly Drosophila melanogaster, however,
the demonstration is straightfor-ward.
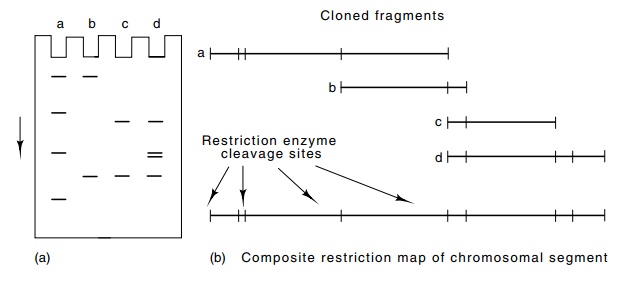
Figure
9.14 (a) A gel showing DNA fragments
resulting from a restrictionenzyme digestion of a set of four partially
overlapping DNA sequences. (b) The four DNAs and their composite.
As explained earlier, the polytene chromosomes in cells of several Drosophila tissues possess characteristic bands that serve as chromosomal landmarks. Therefore the DNA involved in
chromosome rearrange-ments or deletions can readily be observed. Also, highly
radioactive DNA originating from a cloned segment of Drosophila chromosome can be hybridized in situ to Drosophila
polytene chromosomes. The position to which the fragment hybridizes can then be
determined by autoradiog-raphy. Hence the chromosomal position of a cloned
fragment can be determined. If it lies near a gene of interest, walking may be
in order.
A chromosomal walk to clone a specific gene begins
with a restriction map of the cloned piece of DNA. The right- and left-end
terminal restriction fragments are used as probes of a lambda bank. Several
clones are picked that hybridize to the right-hand fragment and not to the
left-hand fragment. Then restriction maps of these clones are made and again
the right- and left-hand fragments are used to find new clones that hybridize
only to the right-hand fragments (Fig. 9.14). The succes-sive lambda
transducing phage that are identified permit walking to the right, and when a
sufficient distance has been covered, in
situ hybridi-zation permits determination of which direction on the cloned
DNA corresponds to moving toward the desired gene.
Related Topics