Chapter: Genetics and Molecular Biology: Genetic Engineering and Recombinant DNA
Plaque and Colony Hybridization for Clone Identification
Plaque and Colony Hybridization for Clone
Identification
Many techniques have been devised for the detection of the desired clones. One of the simplest is genetic selection (Fig. 9.13). Most often this simple avenue is not available, but instead one possesses a related sequence. Sometimes this related sequence is from a gene analogous to the desired gene, but from a different organism. Other times, partial amino acid sequences are known or good guesses can be made as to the amino acid sequence. From such sequences, the DNA sequences can be inferred. In such situations the hybridization ability of complementary DNA sequences can be used to identify cells containing the desired clone.
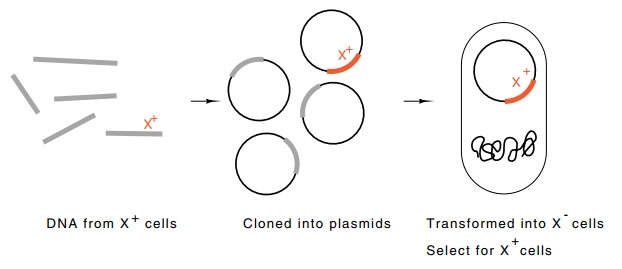
Figure 9.13 Cloning
DNA fromE.coliin which a selection for X+transfor-mants permits
direct selection of the desired clones.
A variety
of direct screening techniques exist for detecting a cloned gene. In these
approaches, cloning the foreign DNA into a lambda vector is convenient because
the phage can accommodate inserted fragments of large size and many lambda
phage may be screened on a single petri plate. The candidate phage collection
is called a lambda bank or library. It is plated out at thousands of plaques
per plate. Then a replica copy of the phage plaques is made by pressing a
filter paper onto the plate. After the paper is removed, it is immersed in
alkali. By these steps the DNA is denatured and fixed onto the paper. Then
radioactively labeled RNA or DNA, called a probe, is hybridized to any
complementary DNA sequences in plaque images on the paper. The probe contains
the known sequence derived from a previously
isolated clone or a sequence de-duced from amino acid sequence information. The
location of the areas with bound probe are then determined by autoradiography,
and viable phage containing the desired insert can be isolated from the
correspond-ing position on the original plate. Analogous techniques exist for
screen-ing colonies containing plasmids.
Related Topics