Chapter: Genetics and Molecular Biology: Genetic Engineering and Recombinant DNA
Plasmid Vectors
Plasmid Vectors
ost
plasmids are small circles that contain the elements necessary for DNA
replication, one or two drug-resistance genes, and a region of DNA into which
foreign DNA may be inserted without damage to essential plasmid functions. One
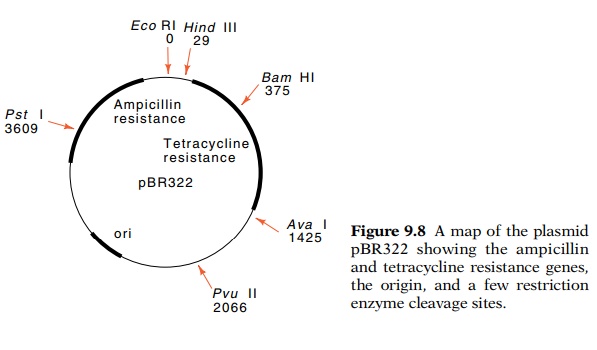
widely used plasmid, pBR322, carries genes coding for resistance to
tetracycline and β-lactamase
(Fig. 9.8). The latter confers resistance to penicillin and related analogs by
cleaving the drugs in the lactam ring, which renders them biologically
inactive. Genes conferring resistance to chloramphenicol, tetracycline, and
A useful
element to have on plasmids is a DNA replication origin from a single-stranded
phage. When such an origin is activated by phage infection, the cell
synthesizes sizeable quantities of just one strand of the plasmid. This
facilitates DNA sequencing. In a typical cloning experiment, a plasmid is cut
in a nonessential region with a restriction enzyme, say EcoRI, foreign EcoRI-cut
DNA is added, and the single-stranded ends are hybridized together and ligated.
Only a small fraction of the plasmids subjected to this treatment will contain
inserted DNA. Most will have recircularized without insertion of foreign DNA.
How can transformants whose plasmids contain inserted DNA be distin-guished
from plasmids without inserted DNA? Of course, in some conditions a genetic
selection can be used to enable only transformants with the desired fragment of
inserted DNA to grow. More often this is not possible and it becomes necessary
to identify candidates that con-tain inserted DNA.
One
method for identifying candidates relies on insertional inactiva-tion of a
drug-resistance gene. For example, within the ampicillin-resis-tance gene in
pBR322 exists the only plasmid cleavage site of the restriction enzyme PstI. Fortunately PstI cleavage generates sticky ends and DNA may readily be ligated
into this site, whereupon it inactivates the ampicillin-resistance gene. The
tetracycline-resistance gene on the plasmid remains intact and can be used for
selection of the cells transformed with the recombinant plasmid. The resulting
colonies can be tested by spotting onto a pair of plates, one containing
ampicillin, and one not containing ampicillin. Only the ampicillin-sensitive,
tetra-cycline-resistant transformants contain foreign DNA in the plasmid. The
ampicillin-resistant transformants derive from plasmid molecules that
recircularized without insertion of foreign DNA.
Another
way to screen for the insertion of foreign DNA utilizes the β-galactosidase gene. Insertion of
foreign DNA within the gene inacti-vates the enzyme, which can be detected by
plating transformed cells on medium that selects for the presence of the
plasmid and also contains substrates of β-galactosidase that produce
colored dyes when hydro-lyzed. A plasmid would be unwieldy if it contained the
complete 3,000 base-pair βgalactosidase
gene. Therefore, only a particular N-terminal portion of the gene is put on the
plasmid. The remainder of the enzyme is encoded by a segment inserted into the
chromosome of the host cells. The two portions of the gene synthesize domains
that bind to one another and yield active enzyme. This unusual phenomenon is
called α-complementation.
Cloning vectors are designed for the insertion offoreign DNA into the short,
N-terminal part of β-galactosidase.
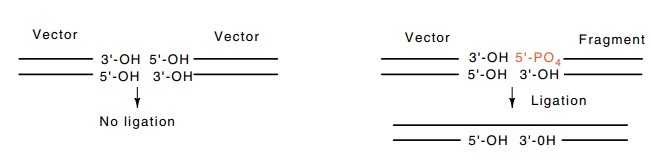
A simple technique can greatly reduce recircularization of vector molecules that lack inserted foreign DNA. If the vector DNA is treated with a phosphatase enzyme after cutting with the restriction enzyme, then recircularization is impossible because the 5’-PO4 ends required by DNA ligase are absent. Foreign DNA, however, will carry 5’-PO4 ends, and therefore two of the four breaks flanking a fragment of foreign DNA can be ligated. This DNA is active in transformation because cells repair the nick remaining at each end of the inserted fragment.
Plasmids or phage cloning vectors can contain a
short stretch of DNA containing unique cleavage sites for a number of
restriction enzymes. These polylinker regions permit cleavage by two enzymes so
that the resulting sticky ends are not self-complementary. The vector cannot
close by itself and be religated; only when a DNA fragment containing the
necessary ends hybridizes to the ends of the plasmid can the structure be
ligated into a closed circle, (Fig. 9.9).
Efficient genetic engineering necessitates that
plasmid DNA be ob-tainable in high quantities. Some plasmids maintain only
three or four copies per cell, whereas other plasmids have cellular copy
numbers of 25 to 50. Many of the high-copy-number plasmids can be further amplified
because the plasmid continues to replicate after protein synthesis and cellular
DNA synthesis have ceased due to high cell densities or the presence of protein
synthesis inhibitors. After amplifi-cation, a cell containing such a
relaxed-control plasmid may contain as many as 3,000 plasmid copies.
Nature did not deliver plasmid vectors ready-made
for genetic engineering. The most useful plasmid vectors were themselves
constructed
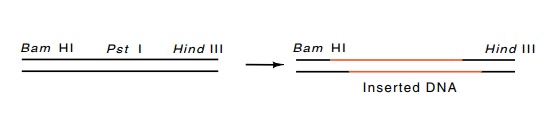
Figure
9.9 Cleavage in a polylinker region
by two different restriction enzymescan create sticky ends that will not
hybridize to themselves, but can only hybridize upon the entry of a foreign DNA
molecule containing the appropriate sticky ends.
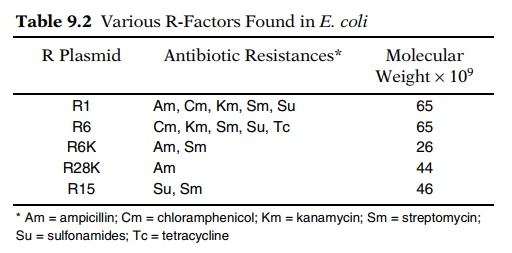
by genetic engineering. The starting materials were
R plasmids, which are plasmids or autonomously replicating DNA elements that
carry one or more drug-resistance genes. The R plasmids are the cause of
serious medical problems, for various bacteria can acquire R plasmids and
thereby become resistant to the normal drugs used for treatment of infections
(Table 9.2). The conversion of an R plasmid into a useful vector requires
elimination of extraneous DNA and removal of multiple restriction enzyme
cleavage sites. In order that foreign DNA can be cloned into the plasmid, the
plasmid should possess only one cleavage site for at least one restriction
enzyme, and this should be in a nones-sential region.
In the construction of cloning vectors, R plasmids
were digested with various restriction enzymes. The resulting mixture of DNA
fragments was hybridized together via the self-complementary ends and then
ligated to produce many combinations of scrambled fragments. This DNA was
transformed into cells. Only new plasmids containing at least the DNA segments
necessary for replication and drug resistance sur-vived and yielded colonies.
The desirable plasmids containing only single cleavage sites for some restriction
enzymes could be identified by amplification and purification of the DNA
followed by test digestions with restriction enzymes and electrophoresis to
characterize the diges-tion products. The plasmid pBR322 possesses single
restriction enzyme cleavage sites for more than twenty enzymes. Some of the
more com-monly used are BamHI, EcoRI, HindIII, PstI, PvuII, and SalI.

Related Topics