Chapter: Pharmaceutical Biotechnology: Fundamentals and Applications : Pharmacokinetics and Pharmacodynamics of Peptide and Protein Drugs
Species Specificity and Allometric Scaling - Pharmacokinetics of Protein Therapeutics
Species Specificity and Allometric Scaling
Peptides and proteins often exhibit distinct species specificity with regard to structure and activity. Peptides and proteins with identical physiological function may have different amino acid sequences in different species and may have no activity or be even immunogenic if used in a different species. The extent of glycosylation and/or sialylation of a protein molecule is another factor of species differences, e.g., for interferon-a or erythropoietin, which may not only alter its efficacy and immunogenicity but also the drug’s clearance. This is of particular importance if the production of human proteins is performed using bacterial cells.
Allometry is a methodology used to relate morphology and body function to the size of an organism. Allometric scaling is an empirical techni-que of predict body functions based on body size. Allometric scaling has found wide application in drug development, especially to predict pharmacokinetic parameters in humans based on the corresponding
parameters in several animal species and the body size differences among these species and humans. Multiple allometric scaling approaches have been described with variable success rates, predominantly during the transition from preclinical to clinical drug development (Dedrick, 1973; Boxenbaum, 1982; Mahmood and Balian, 1999; Mahmood, 2002). In the most frequently used approach, pharmacokinetic parameters between different species are related via body weight using a power function:
P = a Wb
where P is the pharmacokinetic parameter scaled, W is the body weight in kg, a is the allometric coefficient, and b is the allometric exponent. a and b are specific constants for each parameter of a compound. General tendencies for the allometric exponent are 0.75 for rate constants (i.e., clearance, elimination rate constant), 1 for volumes of distribution, and 0.25 for half-lives.
For most traditional small-molecule drugs, allometric scaling is often imprecise, especially if hepatic metabolism is a major elimination pathway and/or if there are interspecies differences in metabolism. For peptides and proteins, however, allometric scaling has frequently proven to be much more precise and reliable, probably because of the similarity in handling peptides and proteins between different mammalian species (Wills and Ferraiolo, 1992). Clearance and volume of distribution of numerous therapeutically used proteins like growth hormone or t-PA follow a well-defined, weight-dependent physiologic relationship between lab animals and humans. This allows relatively precise quantitative predictions for their pharmacokinetic behavior in humans based on preclinical findings (Mordenti et al., 1991).
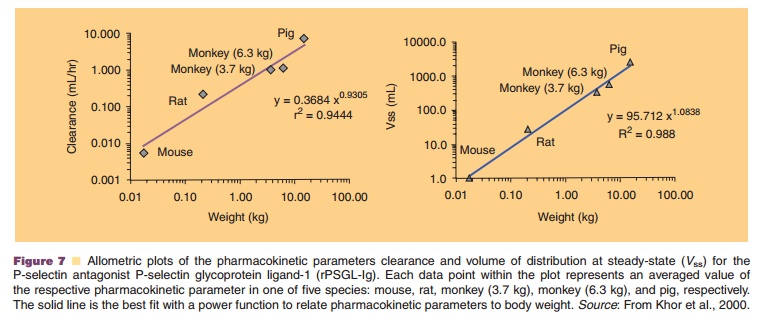
Figure 7, for example, shows allometric plots for the clearance and volume of distribution of a P-selectin antagonist, P-selectin glycoprotein ligand-1 (rPSGL-Ig), for the treatment of P-selectin-mediated diseases such as thrombosis, reperfusion injury, and deep vein thrombo-sis. The protein’s human pharmacokinetic parameters could accurately be predicted using allometric power functions based on data from four species, mouse, rat, monkey and pig (Khor et al., 2000).
It needs to be emphasized that allometric scaling techniques are useful tools for predicting a dose that will assist in the planning of dose-ranging studies, including first-in-man studies, but are not a replace-ment for such studies. The advantage of including such dose prediction in the protocol design of dose-ranging studies is that a smaller number of doses need to be tested before finding the final dose level. Interspecies dose predictions simply narrow the range of doses in the initial pharmacological efficacy studies, the animal toxicology studies, and the human safety and efficacy studies.
Related Topics