Chapter: Pharmaceutical Biotechnology: Fundamentals and Applications : Pharmacokinetics and Pharmacodynamics of Peptide and Protein Drugs
Pharmacodynamics of Protein Therapeutics
PHARMACODYNAMICS OF PROTEIN THERAPEUTICS
Protein therapeutics are usually highly potent com-pounds with steep
dose–effect curves as they are targeted therapies towards a specific,
well-described pharmacologic structure or mechanism. Thus, a care-ful
characterization of the concentration–effect rela-tionship, i.e., the
pharmacodynamics, is especially desirable for protein therapeutics (Tabrizi and
Roskos, 2006). Combination of pharmacodynamics with phar-macokinetics by
integrated pharmacokinetic–pharma-codynamic modeling (PK/PD modeling) adds an
additional level of complexity that allows furthermore characterization of the
dose–exposure–response rela-tionship of a drug and a continuous description of
the time course of effect intensity directly resulting fromthe administration
of a certain dosage regimen (Fig. 9) (Meibohm and Derendorf, 1997; Derendorf
and Meibohm, 1999).
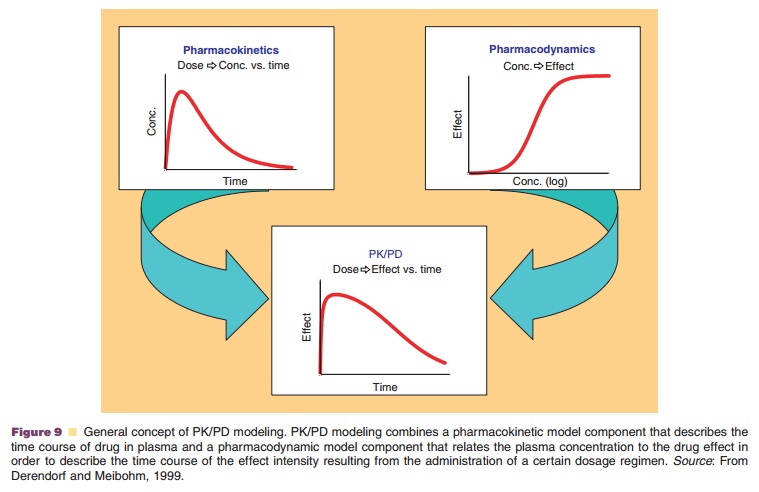
PK/PD modeling is a technique that combines the two classical
pharmacologic disciplines of pharmacoki-netics and pharmacodynamics. It
integrates a pharma-cokinetic and a pharmacodynamic model component into one
set of mathematical expressions that allows the description of the time course
of effect intensity in response to administration of a drug dose. This
so-called integrated PK/PD model allows deriving pharmacoki-netic and
pharmacodynamic model parameters that characterize the dose–concentration–effect
relationship for a specific drug based on measured concentration and effect
data. In addition, it allows simulation of the time course of effect intensity
for dosage regimens of a drug beyond actually measured data, within the
constraints of the validity of the model assumptions for the simulated
condition. Addition of a statistical model component describing inter- and
intraindividual variation in model parameters allows expanding PK/PD models to
describe time courses of effect intensity not only for individual subjects, but
also for whole populations of subjects.
Integrated PK/PD modeling approaches have widely been applied for the characterization of protein therapeutics (Tabrizi and Roskos, 2006). Embedded in a model-based drug development approach, modeling and simulation based on integrated PK/PD does not only provide a comprehensive summary of the avail-able data, but also enables to test competing hypotheses regarding processes altered by the drug, allows making predictions of drug effects under new conditions, and facilitates to estimate inaccessible system variables (Meibohm and Derendorf, 1997; Mager et al., 2003).
Mechanism-based PK/PD modeling appreciat-ing the physiological events
involved in the elabora-tion of the observed effect has been promoted as
superior modeling approach as compared to empiri-cal modeling, especially
because it does not only describe observations but also offers some insight
into the underlying biological processes involved and thus provides flexibility
in extrapolating thetransform this available knowledge into a mechanism-based
PK/PD modeling approach that appropriately char-acterizes the real
physiological process leading to the drug’s therapeutic effect.
The relationship between exposure and response may be either simple or
complex, and thus obvious or hidden. However, if no simple relationship is
obvious, it would be misleading to conclude a priori that no relationship exists
at all rather than that it is not readily apparent (Levy, 1986).
The application of PK/PD modeling is beneficial in all phases of
preclinical and clinical drug develop-ment and has been endorsed by the
pharmaceutical industry, academia and regulatory agencies (Peck et al., 1994;
Breimer and Danhof, 1997; Machado et al., 1999; Lesko et al., 2000; Sheiner and
Steimer, 2000; Meibohm and Derendorf, 2002), most recently by the Critical Path
Initiative of the U.S. Food and Drug Administration (Lesko, 2007). Thus, PK/PD
concepts and model-based drug development play a pivotal role especially in the
drug development process for biologics, and their widespread application
supports a scientifically driven, evidence-based, and focused product
development for protein therapeutics.
While a variety of PK/PD modeling approaches has been employed for
biologics, we will in the following focus on five classes of approaches to
illustrate the challenges and complexities, but also opportunities to
characterize the pharmacodynamics of protein therapeutics:
1. Direct link PK/PD models
2. Indirect link PK/PD models
3. Indirect response PK/PD models
4. Cell lifespan models
5. Complex response models
It should be mentioned, however, that PK/PD models for protein
therapeutics are not only limited to continuous responses as shown in the
following, but are also used for binary or graded responses. Binary responses
are responses with only two outcome levels where a condition is either present
or absence, for example, dead versus alive. Graded or categorical responses
have a set of predefined outcome levels, which may or may not be ordered, for
example the categories “mild,” “moderate,” and “severe” for a disease state.
Lee et al. (2003), for example, used a logistic PK/PD modeling approach to link
cumulative AUC of the anti-TNF-a protein etanercept with a binary
response, the American College of Rheumatology response criterion of 20%
improvement (ARC20) in patients with rheumatoid arthritis.
Related Topics