Chapter: Genetics and Molecular Biology: Transposable Genetic Elements
In vitro Transposition, Threading or Global Topology?
In vitro Transposition, Threading or Global Topology?
Like most systems, deeper understanding of
transposition requires learning the biochemical details of the transposition
reactions. This, in turn, requires assay of the reaction and the use of
purified components. One convenient assay for transposition utilizes a plasmid
containing the DNA sites involved in a transposition reaction. Rearrangement of
segments of the plasmid resulting from transposition will change the sizes
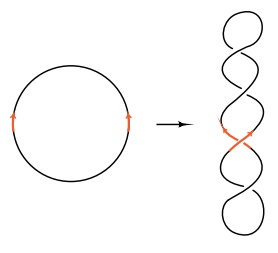
Figure
19.13 Well-separated in-verted repeats
of a sequence on a circle can be adjacent to one an-other and parallel when the
circle is supercoiled.
Experiments with a plasmid containing two res sites of Tn3 showed that resolution
would proceed in vivo in the presence
of a Tn3 with a functional resolvase gene. The reaction would also take place in vitro upon the addition of purified
resolvase. This reaction occurred, how-ever, only if the plasmid DNA was
supercoiled and if the two res sites
on the plasmid were correctly oriented. Reversing one blocked the reaction, in
contrast to what is seen with lambda phage where the excision reaction will
proceed with the att sites in either
orientation with respect to one another.
Why should an orientation requirement exist? We
might expect that the two sites encounter each other by random flailings about
of the DNA, in which case, there could be no orientation requirement. One
attractive hypothesis to explain the orientation requirement is threading. Once
the protein binds one site, it reaches the other by crawling along the DNA,
thus preserving its knowledge of the orientation of the first site. Thread-ing
the DNA through the protein until the second site arrives at the protein
accomplishes the same end.
The requirement for a specific orientation for the
transposition sites can be explained in a different way. Suppose that the sites
had to interact in a specific orientation at the time of transposition. Due to
negative supercoiling, two sites which are in inverted repeat orientation on a
circular DNA molecule will interact in a parallel orientation (Fig. 19.13).
Interacting in an antiparallel orientation would require massive rear-rangement
of the supercoils and would be highly unlikely to occur.
The two possible explanations for the orientation
requirement of the interacting sites of the Mu-Tn3 class of transposons can be
tested by making use of the lambda phage integration enzymes. After a plasmid
containing Mu sites which are flanked by lambda phage attachment sites has
supercoiled itself, then reaction between the lambda att sites can be performed. Such an interaction should have little
effect upon the overall supercoiled structure of the plasmid. Yet, if the sites
are in one
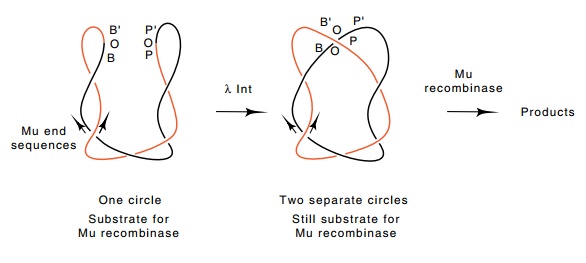
Figure
19.14 Action of lambda phage Int
protein on a circle containing the Muends can separate it into two chemically
separate circles without dramatically altering the wrapping structure in the
vicinity of the Mu ends.
orientation, the reaction will invert the Mu sites,
and if the lambda att sites are in
the opposite orientation, the reaction will generate two separate DNA
molecules, each containing one of the Mu sites (Fig. 19.14). Neither
orientation of the lambda att sites
had any effect on the Mu transposition reaction. This result shows that
tracking is not used by the Mu enzymes to determine the transposition sites’
orientations, but rather it is the orientation of the sites with respect to one
another in the supercoiled DNA that matters.
Related Topics