Chapter: Genetics and Molecular Biology: Transposable Genetic Elements
IS Elements in Bacteria
IS Elements in Bacteria
Nonsense mutations in many operons have polar effects, that is, they reduce the expression of downstream genes. In efforts to study these effects more carefully, in the late 1960s Malamy, Shapiro, and Starlinger each isolated and characterized strongly polar mutations in the lac and gal operons. In addition to the expected classes of nonsense mutations,another type was also found. This new type was highly polar but not suppressible by known nonsense suppressors. Since the mutations reverted, albeit at very low rates, it seemed likely that they were inser-tions and not deletions. Therefore it became of great interest to find out just what these mutations were. Nowadays this question would probably be answered by cloning a portion of the DNA containing the mutation and sequencing it to determine the exact nature of the mutation. Such a procedure was impossible at the time and less direct experiments were performed.
One likely cause of the polarity was the insertion
of extraneous nucleotides into the lac
and gal genes. Fortunately, this
possibility could be tested by the techniques then available. If the insertions
were larger than a few hundred nucleotides, they could be detected directly
because they would increase the density of lac
or gal transducing phage. There-fore
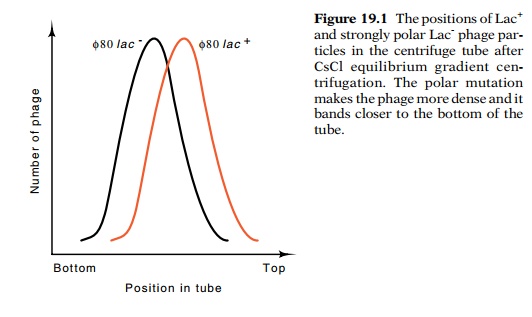
the mutations were crossed onto λdgal and φ80plac
phage and the densities of phage both with and without the mutations were
carefully compared by equilibrium gradient centrifugation (Fig. 19.1). The
results showed that transducing phage carrying the polar mutations were denser
than the corresponding transducing phage not carrying the mutations. Therefore
the mutations were caused by the insertion of DNA elements. These later came to
be called IS, for insertion sequences.
At first, the insertions were suspected of being
merely random se-quences that somehow had been inserted into the genes being
studied. The mechanism of their generation is not so easy to imagine, however.
Point mutations, insertions and deletions of a nucleotide or two, and sizable
deletions may all be created by simple reactions likely to occur in cells. Not
so for sizable insertions.
Additional study of the IS sequences began to
clarify their origin. One particularly valuable technique for their study was
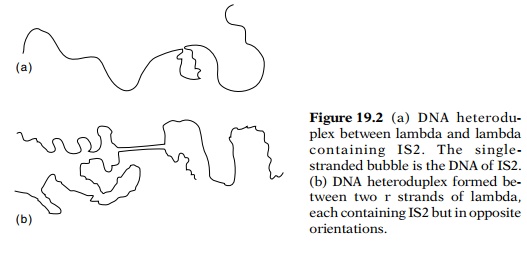
the formation of DNA heteroduplexes (Fig. 19.2). DNA isolated from the gal and lac transducing phage with and without the IS sequences was used in
these experiments. When necessary, the strands of these phage could even be
separated and purified. The lengths of IS sequences fell into discrete size
classes, and any particular IS sequence was found to insert into a gene in
either of the two possible orientations. The existence of the length classes
suggested that a small number of IS sequences might be responsible for all the
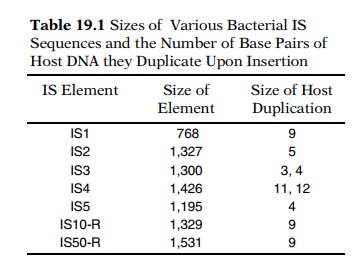
insertions. Indeed, fewer than ten different IS sequences have been found in Escherichia coli (Table 19.1). The demon-stration
that IS elements were specific sequences did not explain how they were inserted
into different sites in the cell’s chromosome, but by analogy to lambda phage,
the elements were expected to code for one or more proteins involved in the
process. By now the IS elements have been obtained on phage and plasmids and
they have been sequenced and characterized using in vitro mutagenesis as described below. From these studies a
comprehensive picture of their structure and function has emerged.
Related Topics