Chapter: Genetics and Molecular Biology: Genetics
Fate Mapping and Study of Tissue-Specific Gene Expression
Fate Mapping and Study of Tissue-Specific Gene
Expression
One obvious way to examine the tissue specificity
of gene expression is to isolate the tissues and assay each for the protein or
gene product in question. A slight modification of this approach is quite
reasonable. The synthesis of messenger from the various tissues of a fly can be
measured approximately by DNA-RNA hybridization. As we shall see in a later,
DNA from desired genes can be obtained and then used in such in situ hybridization experiments.
Remarkably, genetics experimentscalled fate mapping can also locate the tissues
in which an altered gene is expressed. This approach does not require knowledge
of the gene involved and therefore it is useful in initial steps of studies.
Genetic engineering has also developed techniques for examination of tissue-specific
gene expression, but those approaches first require isolating DNA or RNA of the
gene in question. Fate mapping is useful when the gene involved is unknown. As
an example of localizing the activity of a gene, imagine a mutant fly that is unable
to flap its wings. This could be because of a defective wing, a defective wing
muscle, a defective nerve to the muscle, or defective neurons in the brain.
Fate mapping can determine which tissue is responsible for such altered
behavior.
Fate mapping relies on the developmental pathway of
the fly. A fertilized Drosophila egg
contains a single cell whose nucleus undergoes about nine divisions. These
nuclei then migrate to the surface of the egg to form the blastula stage, and
three more divisions occur before cell walls are laid down. At this stage,
different cells on the surface ultimately
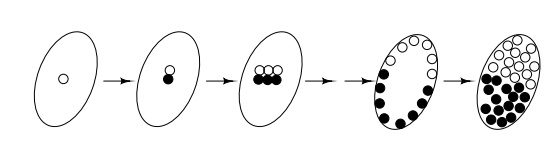
become different parts of the adult fly, but cells
lying near to each other frequently develop into adjacent parts on the fly.
Therefore, a map can be drawn on the egg of the parts of the adult fly that
each of these cells will become. If it were possible to associate a particular
phenotype in an adult with a particular location on the egg, then the tissue
responsible for the adult phenotype would be determined. This is possible!
The association of tissues in the mature fly with
positions on the blastula utilizes selective chromosome loss during development
of the blastula. Fly development is not greatly altered in a female egg cell if
one of the X chromosomes contains a defect so that it begins the first nuclear
replication a little late. Consequently, this chromosome often is not
segregated into one of the two resultant daughter nuclei resulting from the
first nuclear division in the egg. The final result of this chromosome loss is
that about half of the cells of the blastula will be diploid XX and the others
will be haploid X. Since the spatial orientation of the first cleavage with
respect to the egg shell is not uniform from egg to egg, and because there is
little mixing of the nuclei or cells during subsequent divisions, different
sets of cells will be XX and X in different blastulas. Suppose that these two
types of cells can be distinguished. This can be done by placing a recessive
body color marker gene, for example yellow, on the stable X chromosome. Then
cells of the fly possessing the XX genotype will be black and the X genotype
will be yellow. The adult fly will have a mottled appearance.
The probability that two different body parts
possess different colors will be proportional to the distance that their
corresponding ancestor cells in the blastula state were separated. The greater
their separation, the greater the chances that the line separating the two cell-types
will fall between them. If they are close together, there is little chance they
will be of different cell-type and therefore there is little chance they will
possess different body color.
A collage appears when the body parts of the adult
fly are mapped to the blastula. The map can then be used as follows to locate
the tissue in which a recessive mutation is expressed. If the mutation is
located on the X chromosome which is not lost during the development, then
those tissues which can express the mutant phenotype will be haploid. For
example, if the mutant phenotype always appears in flies with haploid second
left legs and only appears in these flies, then it can be concluded that the
tissue in which the mutation is expressed is the second left leg. More
generally however, the frequency of association of the mutant phenotype with a
number of landmarks gives the distance on the blastula between the landmarks
and the tissue in question. Transferring these distances to the blastula fate
map then reveals the tissue in question.
Related Topics