Chapter: Genetics and Molecular Biology: Genetics
Elements of Recombination in E. coli, RecA, RecBCD, and Chi
Elements of Recombination in E. coli, RecA, RecBCD, and Chi
What types of biochemical evidence can be found in
support of the mechanisms of recombination discussed in the previous section?
His-torically, the elucidation of metabolic pathways was often assisted by the
isolation of mutations in enzymes catalyzing individual steps of the pathway.
With similar objectives in mind, mutations were isolated that decreased or
increased the ability of phage, E. coli, yeast, and some other organisms to
undergo recombination. Subsequently, the enzyme prod-ucts of the recombination
genes from E. coli and yeast have been identified and purified. Of greatest
importance to recombination are the RecA and RecBCD proteins. Additional enzyme
activities that might be expected to play roles, such as DNA ligase, DNA
polymerases, single-stranded binding protein, and proteins that wind or unwind
DNA, have already been discussed and will not be further mentioned here.
RecA mutants are unable to engage in genetic
recombination, and asexpected, the purified RecA protein possesses a variety of
activities that appear related to recombination. In the presence of ATP the
protein binds to single-stranded DNA in a highly cooperative manner. Once one
molecule has bound to a stretch of single-stranded DNA, other mole
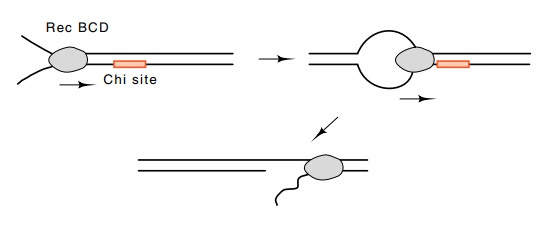
Figure
8.16 RecBCD binds to a free end of the
DNA and moves at more than200 base pairs per second. As RecBCD crosses a chi
site, one of the strands is cut.
The known activities of the RecBCD protein suggest
that this protein generates the single-stranded regions that initiate the
strand invasion process catalyzed by RecA. RecBCD protein binds at the free end
of a double-stranded DNA duplex and moves down the DNA separating the DNA
strands and leaving a single-stranded region in its wake. If it encounters a
specific eight nucleotide sequence called a chi
sequence, the enzyme cleaves one of the strands, leaving a single-stranded DNA
end that can be covered with RecA protein and initiate recombination (Fig.
8.16).
Chi can stimulate recombination at distances up to
10,000 bases away from itself and appears to act only in one direction from
itself, that is, it possesses a polarity. Only when chi is crossed by RecBCD traveling in one direction does cleavage
occur. Apparently organisms other than bacteria utilize sequences that function
like chi, since hotspots of genetic
recombination are found in yeast and fungi as well. The fact that RecBCD can
enter DNA only at an end restricts its stimulation of recombination to those
situations in which ends are generated. In addition to random chromosome
breakage, in which case genetic re-combination may be a good way to attempt
rescue, ends are generated during transfer of genetic information. The rate of
genetic recombina-tion between homologous DNA sequences during normal growth is
very low. Recombination primarily occurs at the time of introduction of new DNA
into bacteria. The analogous principle applies to meiosis of eu-karyotic cells,
but the recombination stimulating process there is not yet known.
Related Topics