Chapter: Genetics and Molecular Biology: Genetics
Point Mutations, Deletions, Insertions, and Damage
Point Mutations, Deletions, Insertions, and
Damage
The structure of DNA permits only three basic types
of alteration or mutation at a site: the substitution of one nucleotide for
another, the deletion of one or more nucleotides, and the insertion of one or
more nucleotides. A nucleotide substitution at a point is called a transition
if one purine is substituted for the other or one pyrimidine is substituted for
the other and is called a transversion if a purine is substituted for a
pyrimidine or vice versa
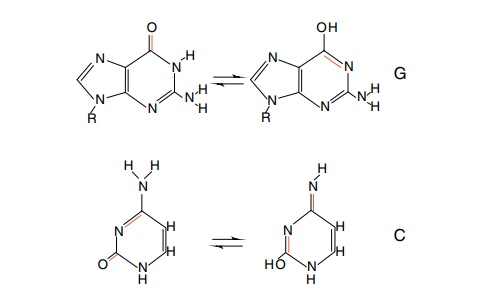
Figure
8.1 Tautomeric forms of guanine and
cytosine base pair differently dueto alternations of hydrogen bond donating and
accepting groups.
In addition to substitutions of one nucleotide for
another in single-stranded DNA or one base pair for another in double-stranded
DNA, nucleotides are susceptible to many types of chemical modification. These
can include tautomerizations and deamination or more extensive damage such as
the complete loss of a base from the ribose phosphate backbone (Fig. 8.1). The
cellular repair mechanisms, however, remove many such modified bases so that
the gap can be refilled with normal nucleotides. Those modified bases that
escape repair cannot themselves be passed on to the next generation because, on
DNA replication, one of the usual four nucleotides is incorporated into the
daughter strand opposite the altered base. Frequently the base so incorporated
will be incorrect, and consequently, a mutation is introduced at such a
position.
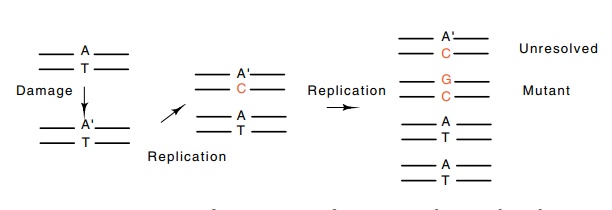
Mutations arise from a variety of sources. As
discussed, point mutations can occur spontaneously during replication of the
DNA through the misincorporation of a nucleotide and the failure of the editing
mechanisms to correct the mistake or through the chemical instability of the
nucleotides. For example, cytosine can deaminate to form uracil, which is then
recognized as thymine during DNA replica-tion.
The frequency of spontaneous appearance of point
mutations often is too low for convenient experimentation, and mutagens are
therefore used to increase the frequency of mutants in cultures 10 to 1,000
times above the spontaneous frequency. A variety of mutagens have been
discovered, some by rational considerations and some by chance. Many are
nucleotide analogs that are incorporated into the DNA instead of the normal
nucleotides. These increase the frequency of mispairing in subsequent rounds of
DNA replication. Other mutagens are chemically reactive molecules that damage
or modify bases in DNA. Ultraviolet light is also a mutagen as discussed
earlier. The damage it creates ultimately leads to mutations either through a
reduced fidelity of syn-thesis or an elevated probability of mistaken repair of
the original damage. In one way or another, mutagens increase the frequency of
generating mispaired bases or increase the frequency that mispaired bases
escape repair. Ultimately, either leads to an increase in the probability that
the original DNA sequence will be changed.
The mechanisms generating deletions and insertions
are not as well understood. Errors in DNA replication provide plausible
mechanisms for the generation of one or two base insertions or deletions. Most
likely slippage, perhaps stimulated by an appropriate sequence, will permit a
Figure
8.2 Two mechanisms for generating
deletions between repeated sequences. The first is looping with recombination
between points within a single chromosome, and the second is unequal crossing
over between two chromosomes.
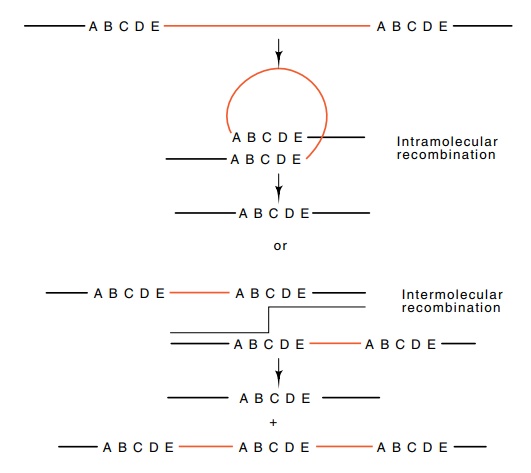
Insertions and deletions larger than a few bases
arise by a different mechanism. The end points of a number of deletions in
bacteria are located at short, repeated or almost repeated sequences. The
deletion removes one of the repeats and the intervening sequence. We can
picture two plausible events that could create such deletions (Fig. 8.2). The
first is a looping of a single chromosome followed by elimination of the
material between the two repeats. The second is similar to the first, but it
occurs between two chromosomes and transfers the material from one chromosome
to the other. One chromosome suffers a deletion and the other an insertion.
The creation of deletions is also stimulated by the
presence of some genetic elements called insertion sequences or transposons.
These ele-ments transpose themselves or copies of themselves into other sites
on the chromosome. In the process they often generate deletions in their
vicinity.
Related Topics