Chapter: Essential Microbiology: Microbial Metabolism
Enzymes
Enzymes
An enzyme is a cellular catalyst; it makes biochemical reactions proceed many times more rapidly than they would if uncatalysed. The participation of an enzyme can in-crease the rate of a reaction by a factor of millions, or even billions.
Traditionally, all enzymes have been thought of as globular proteins, but around twenty years ago it was demonstrated (surprisingly) that certain RNA molecules also have catalytic properties. These ribozymes however, are very much in the minority, carrying out specific cut-and-splice reactions on RNA molecules, and in the present
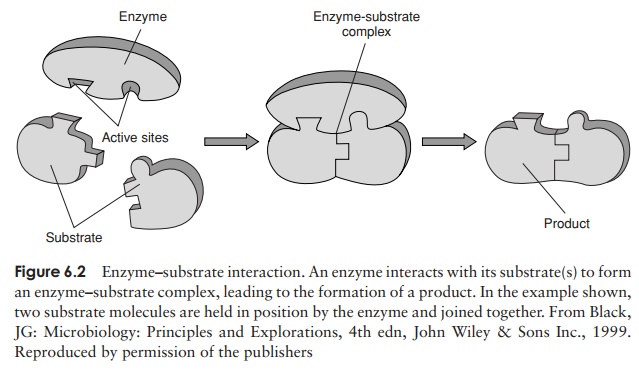
Like any other catalyst, an enzyme remains unchanged at the end of a reaction. It must, however, at some point during the reaction bind to its substrate (the substance upon which it acts) to form an enzyme–substrate complex (Figure 6.2) by multiple weak forces such as electrostatic forces and hydrogen bonding. Only a small part of the enzyme’s three-dimensional structure is involved in this binding; these few amino acids make up the active site, which forms a groove or dent in the enzyme’s surface, into which the appropriate part of the substrate molecule fits (Figure 6.3). The amino
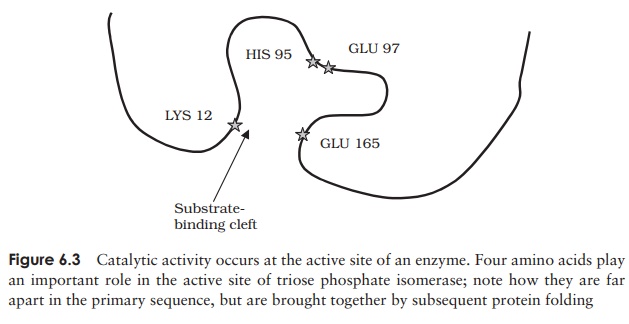
acid residues that go to make up the active site may be widely separated in the enzyme’s primary structure, but by means of the secondary and tertiary folding of the molecule, they are brought together to give a specific conformation, complementary to that of the substrate. It is this precise formation of the active site that accounts for one of the major characteristics of enzymes, their specificity. You should not think, however, that these few residues making up the active site are the only ones that matter; the enzyme can only fold in this way because the order and arrangement of the other amino acids allows it.
Enzyme classification
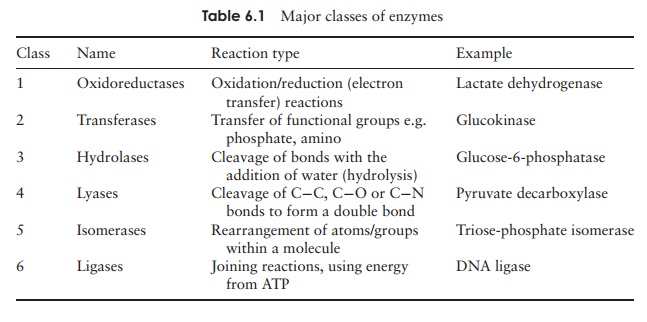
Most enzymes have names that end in the suffix –ase. The first part of the name often gives an indication of the substrate; for example, urease and pyruvate decar-boxylase. Other enzymes have names that are less helpful, such as trypsin, and others have several alternative names to confuse the issue further. To resolve such problems, an internationally agreed system of nomenclature has been devised. All enzymes are assigned initially to one of six broad groups according to the type of reaction they carry out, as shown in Table 6.1. Each enzyme is then placed into successively more specific groupings, each with a number. Thus regardless of any colloquial or alternative names, each enzyme has its own unique and unambiguous four-figure Enzyme Commission ‘signature’ (pyruvate decarboxylase, mentioned above, is EC 4.1.1.1).
Certain enzymes have a non-protein component
Many enzymes require the involvement of an additional, non-protein component in order to carry out their catalytic action. These ‘extra’ parts are called cofactors; they
Some of the most important coenzymes act by transferring electrons between substrate and product in redox reactions .
How do enzymes speed up a reaction?
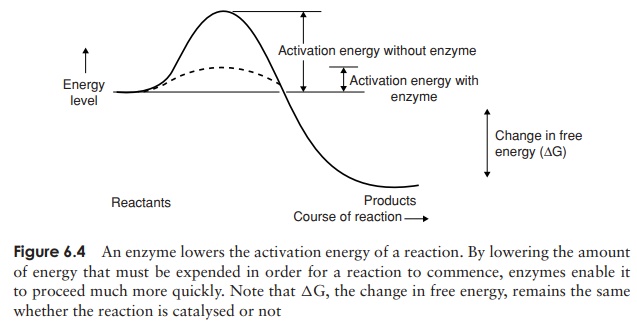
For any chemical reaction to take place there must be a small input of energy. This is called the activation energy,and is often likened to the small push that is needed to loosen a boulder and allow it to roll down a hill. It is the energy needed to convert the molecules at the start of a reaction into intermediate forms known as transition states, by the rearrangement of chemical bonds. The great gift of enzymes is that they can greatly lower the activationenergy of a reaction, so that it requires a smaller energy input, and may therefore occurmore readily (Figure 6.4).
Environmental factors affect enzyme activity
The rate at which an enzyme converts its substrate into product is called its velocity (v), and is affected by a variety of factors.
Temperature
The rate of any chemical reaction increases with an increase in temperature due to the more rapid movement of molecules, and so it is with enzyme-catalysed reactions, until a peak is reached (the optimum temperature) after which the rate rapidly falls away. What
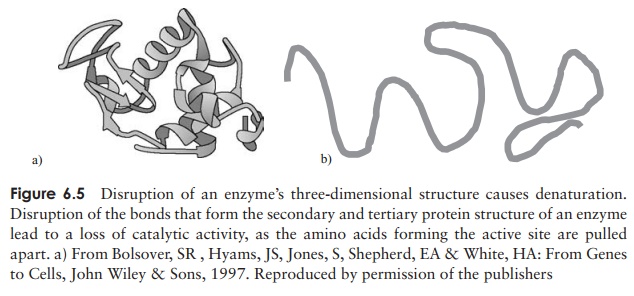
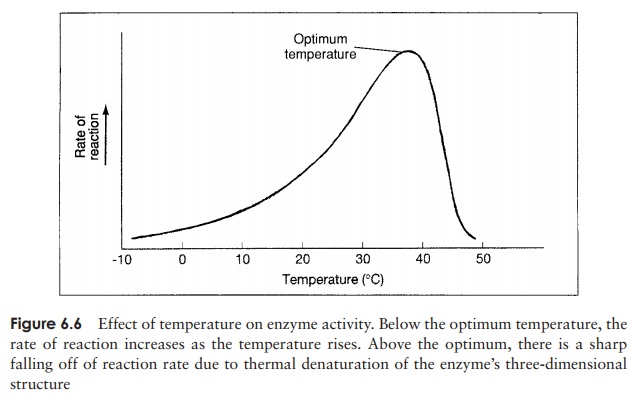
causes this drop in the velocity? Recall that the very ordered secondary and tertiary structure of a protein molecule is due to the existence of numerous weak molecular bonds, such as hydrogen bonds. Disruption of these by excessive heat results in denaturation (Figure 6.5), that is, an unfolding of the three-dimensional structure. In the case of an enzyme, this leads to changes in the configuration of the active site, and a loss of catalytic properties. The effect of temperature on enzyme activity is shown in Figure 6.6. The graph can be thought of as a composite of two lines, one increasing with temperature due to the rise in thermal energy of the substrate molecules, and one falling due to denaturation of the protein structure. Before the optimum temperature it is the former which dominates, then the effect of the latter becomes more pronounced, and takes over completely.
pH
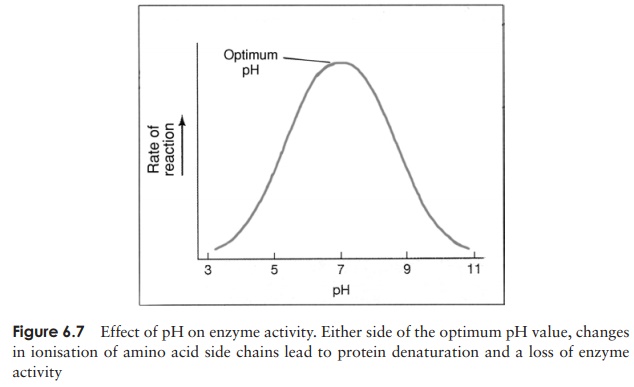
Enzyme velocity is similarly affected by the prevailing pH. Once again, this is due to alterations in three-dimensional protein structure. Changes in the pH affect the ioni-sation of charged ‘R’-groups on amino acids at the active site and elsewhere, causing changes in the enzyme’s precise shape, and a reduction in catalytic properties. As with temperature, enzymes have an optimum value at which they operate most effectively; when the pH deviates appreciably from this in either direction, denaturation occurs, leading to a reduction of enzyme activity (Figure 6.7).
Microorganisms are able to operate in a variety of physicochemical environments, a fact reflected in the diversity of optimum values of temperature and pH encountered in their enzymes.
Substrate concentration
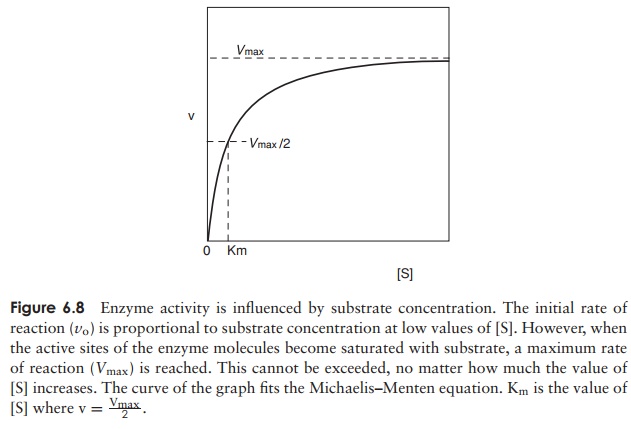
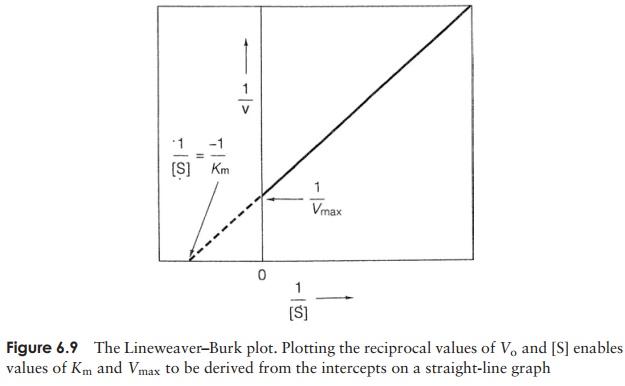
Under conditions where the active sites of an enzyme population are not saturated, an increase in substrate concentration will be reflected in a proportional rise in the rate of reaction. A point isreached, however, when the addition of further sub-strate has no effect on the rate (Figure 6.8). This is be-cause all the active sites have been occupied and theenzymes are working flat out; this is called the max-imum velocity (Vmax). A measure of the affinity an en-zyme has for its substrate (i.e. how tightly it binds to it) isgiven by its Michaelis constant (Km). This is the substrate concentration at which the rate of reaction is half of the Vmax value. Values of Vmax and Kmare more easily determined experimentally by plotting the reciprocals of [S] and V to obtain a straight line (Figure 6.9).
Some enzymes do not obey Michaelis–Menten kinetics. The activity of allostericenzymes is regulated by effector molecules which bind at a position separate from theactive site. By doing so, they induce a conformational change in the active site that results in activation or inhibition of the enzyme. Thus effector molecules may be of two types, activators or inhibitors.
Enzyme inhibitors
Many substances are able to interfere with an enzyme’s ability to catalyse a reaction. As we shall see, enzyme inhibition forms the basis of several methods of microbial control, so a consideration of the main types of inhibitor is appropriate here.
Perhaps the easiest form of enzyme inhibition to understand is competitive inhibition. Here, the inhibitory substance competes with the normal substrate for access to the enzyme’s active site; if the active site is occupied by a molecule of inhibitor, it can’t bind a molecule of substrate, thus the reaction will proceed less quickly (Figure 6.10a).
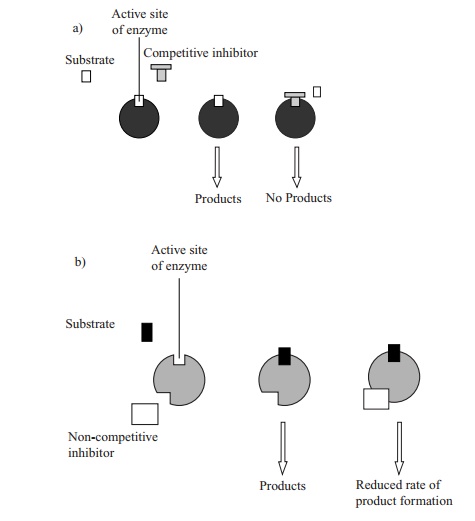

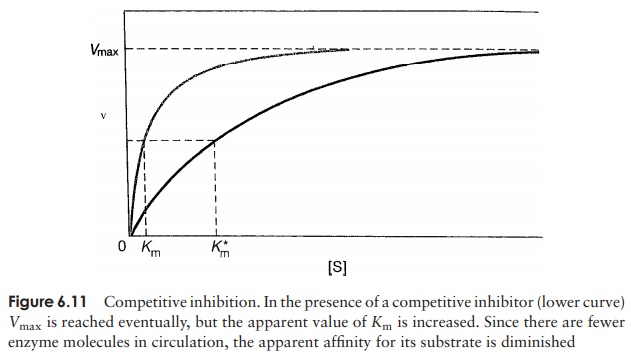
The competitive inhibitor is able to act in this way because its molecular structure is sufficiently similar to that of the substrate for it to be able to fit into the active site. The effect is competitive because it depends on the relative concentrations of the substrate and inhibitor. If the inhibitor is only present at a low concentration, its effect will be minimal, since the number of enzyme–inhibitor interactions will be greatly outweighed by reactions with the ‘proper’ substrate. The Vmax value for the enzyme is not reduced, but it is only reached more gradually. The apparent affinity of the enzyme for its substrate is decreased, reflected by an increase in the Km (Figure 6.11).
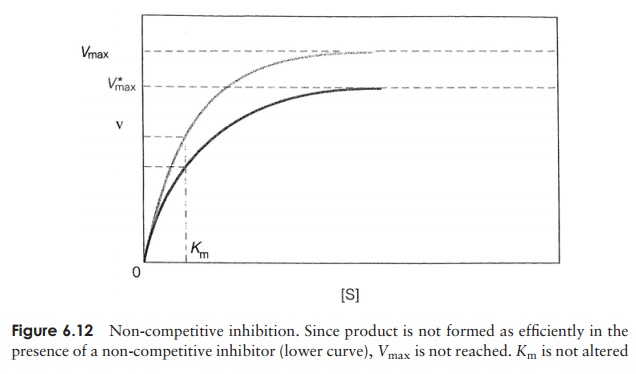
Not all inhibitors act by competing for the active site, however. Non-competitiveinhibitors act by binding to a different part of the enzyme and in so doing alter itsthree-dimensional configuration (Figure 6.10b). Although they do not affect substrate binding, they do reduce the rate at which product is formed. Vmax cannot be reached; however, the value of Km, is unchanged (Figure 6.12). Such inhibitors may bind to either the enzyme–substrate complex or to free enzyme. Both competitive and non-competitive forms of inhibition are reversible, since the inhibitor molecule is relatively weakly bound and can be displaced.
Irreversible inhibition, on the other hand, is due to the formation of a strong covalentlinkage between the inhibitor and an amino acid residue on the enzyme. As a result of its binding, the inhibitor effectively makes a certain percentage of the enzyme population permanently unavailable to catalyse substrate conversion.
Related Topics