Chapter: Genetics and Molecular Biology: Transcription,Termination, and RNA Processing
The Discovery of Self-Splicing RNAs
The Discovery of Self-Splicing RNAs
Cech found that the nuclear ribosomal RNA from Tetrahymena contains an intervening
sequence. In efforts to build an in vitro
system in which splicing would occur, he placed unspliced rRNA in reactions
containing and lacking extracts prepared from cells. Amazingly, the control
reac-tions for the splicing reactions, those lacking the added extract, also
spliced out the intervening sequence. Naturally, contamination was suspected,
and strenuous efforts were made to remove any possible Tetrahymena proteins from the substrate RNA, but splicing in
theabsence of Tetrahymena extract
persisted. Finally, Cech placed the gene for the rRNA on a plasmid that could
replicate in E. coli, prepared the DNA from E.
coli cells, a situation that had to
be devoid of any hypothetical Tetrahymena
protein, and he still found splicing. This proved, to even the most skeptical,
that the Tetrahymena rRNA was
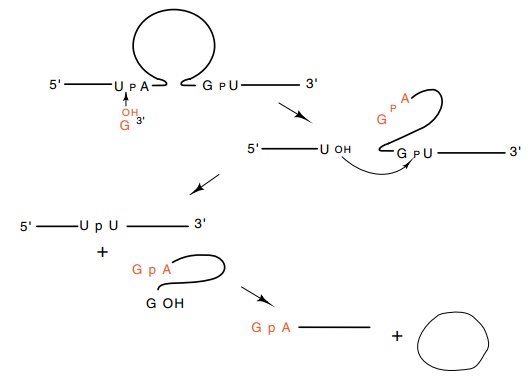
Figure
5.20 The series of self-splicing
reactions followed byTetrahymenarRNA.
As shown in Fig. 5.20, guanosine is essential to
this self-splicing reaction, but it does not contribute chemical energy to the
splicing products. Further studies on the Tetrahymena
self-splicing reaction show that not only is a 480 nucleotide section of RNA
removed from the middle of the ribosomal RNA, but the removed portion then goes
on to close on itself forming a circle and releasing a short linear fragment.
At first it seems surprising that neither ATP nor any other energy source is
required for the cutting and splicing reactions. The reason is that external
energy is not required. Chemically, all the reactions are tran-sesterifications
and the numbers of phosphodiester bonds are con-served. One might ask, then,
why the reaction proceeds at all. One answer is that in some cases the products
of the reactions are three polynucleotides where initially only one plus
guanosine existed before. Together these possess higher entropy than the
starting molecules, and therefore their creation drives the reaction forward.
The transesterification reactions involved in the
self-splicing proceed at rates many orders of magnitude faster than they could
with ordinary transesterifications. Only two reasons can explain the stimulated
rate of these reactions. First, the secondary structure of the molecules can
hold the reactive groups immediately adjacent to one another. This increases
their effective collision frequencies far above their normal solution values.
The second reason is that the probability of a reaction occurring with a
collision can be greatly improved if the bonds involved are strained. Studies
with very small self–cutting RNAs and also molecular dynamics calculations
indicate that such a strain is crucial to the reactions. Undoubtedly,
self–splicing utilizes both principles.
Self–splicing has been found in two of the
messenger RNAs of the bacteriophage T4, and in the splicing of mRNA in the
mitochondrion of yeast. The mitochondrial self–splicing introns comprise a
second group of self–splicing introns. Their secondary structure differs from
those of the Group I self–splicing introns of which the Tetrahymena rRNA intron is an example. The Group II introns do not
use a free guanosine to initiate the splicing process. They use an internal
nucleotide, and in this respect use a reaction mechanism more like that used in
processing pre–mRNA. The existence of splicing in bacteria and eukaryotes
sug-gests that splicing in general has existed in the common precursor to both
organisms. The scarcity splicing events in prokaryotes might be a result of the
greater number of generations they have had in which to select for the loss of
introns. Eukaryotes may still be struggling with the “infection.”
Related Topics