Chapter: Genetics and Molecular Biology: Transcription,Termination, and RNA Processing
Other RNA Processing Reactions
Other RNA Processing Reactions
The precursor tRNAs in yeast contain an intervening
sequence. This is removed in a more traditional set of enzymatic cleavage and
ligation reactions (Fig. 5.23).
Plant viruses frequently have RNA genomes. These
viruses can them-selves have viruses. These are known as virusoids, and they
can grow in cells only in the presence of a parental virus. Virusoids do not
encode any proteins, but they are replicated. Part of their replication cycle
requires the specific cleavage of their RNA molecules. This they do in a
self–cutting reaction. Symons and Uhlenbeck have investigated the
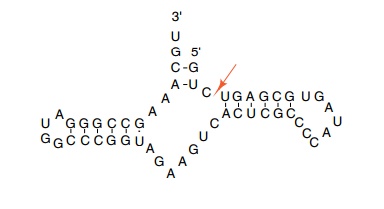
minimal nucleotide requirements for self–cleavage
of these molecules. Remarkably, it is a scant 25 nucleotides that can form into
a functional
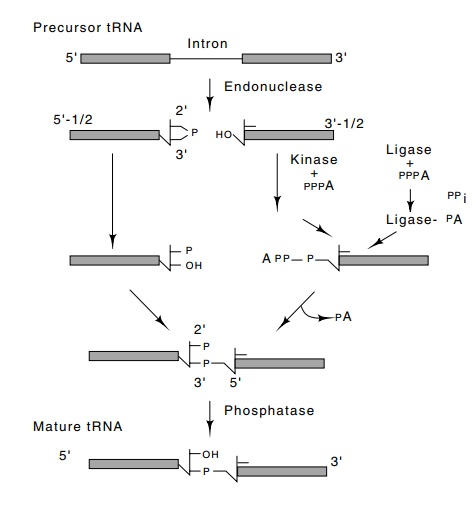
Figure
5.23 The pathwayfor cutting and
splicing tRNA followed by the yeast Sac-charomyces.
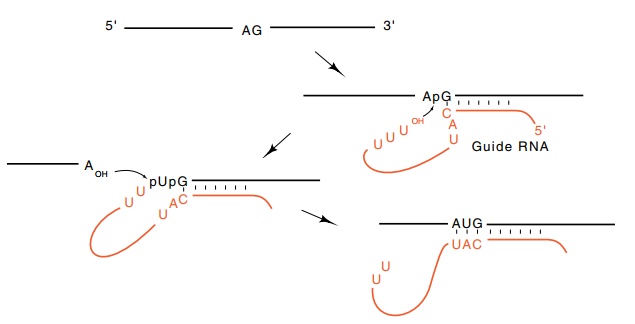
Figure
5.24 Editing of pre-mRNA by a guide
RNA using transesterifications.
Simple editing of a nucleotide or two has been
observed in a mammalian RNA, but in the mitochondrion of some protozoa far more
extensive editing has been observed. This raises the question of just where the
informa-tion for the editing is stored. Changing a single base can conceivably
be a result of a set of reactions catalyzed by enzymes designed for just the
sequence at which the change occurs, but in the more dramatic exam-ples of
editing, in which more than 50 U’s are inserted to produce the final edited
sequences, far too many different enzymes would be re-quired. Initial
examinations of the DNA of the organisms, both with computer searches of known
sequence and hybridization studies, failed to reveal any sequences that could
have encoded the edited sequence. Eventually it was found that the information
for the edited sequences was carried in short RNAs complementary to segments of
the final edited sequence. These are called guide sequences. Although cutting
and re-ligation could be the pathway for editing, intermediates in editing are
found that indicate instead, that the guide sequences transfer U’s from their
3’ ends to the necessary positions in the mRNA by transesterifica-tion
reactions analogous to those used in splicing (Fig. 5.24).
Related Topics