Chapter: Genetics and Molecular Biology: Transcription,Termination, and RNA Processing
Processing Prokaryotic RNAs After Synthesis
Processing Prokaryotic RNAs After Synthesis
It is not altogether surprising that intact
transcripts produced by tran-scription of some operons are not always suited
for all the biological roles of RNA. In E.
coli the ribosomal and tRNAs are
processed following transcription. For example, RNAse III first cleaves in the
portions of the rRNA that are folded back on themselves to form hairpins (Fig.
5.8). Then other nucleases make additional cuts. RNAse III also cleaves the
early transcripts of phage T7. A number of methyl groups are also added to the
rRNA after its synthesis.
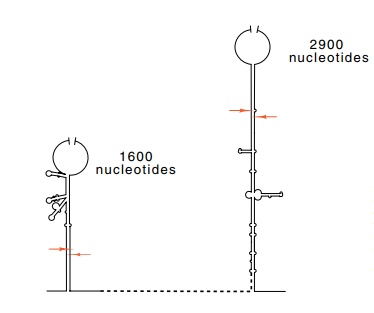
Figure 5.8 Schematic of the basepaired stems surrounding the 16S and 23S ribosomal RNAs. The ar-rows mark the points of RNAse III cleavage.
How do we know RNAse III cleaves the ribosomal and
phage T7 RNAs? The effects of mutations are one way to attack this general
problem of demonstrating the in vivo
role of an enzyme. Investigators first isolated a mutant defective in RNAse
III. They did this by screening extracts prepared from isolated colonies of
mutagenized cells for the activity of the enzyme RNAse III (Fig. 5.9). One out
of 1,000 colonies yielded cells with greatly depressed levels of the enzyme.
Subsequently,
Figure
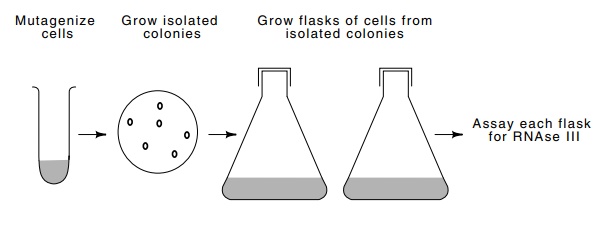
5.9 An isolation scheme for RNAse III
mutants. Mutagenized cells arediluted and spread on plates so as to yield
colonies derived from individual cells. Cultures are grown from these colonies
and then tested for the presence of RNAse III.
these cells were found to possess appreciable
quantities of uncleaved ribosomal RNA. Furthermore, this RNA could be cleaved in vitro with purified RNAse III to
yield the smaller ribosomal RNAs.
Although it occurs very rarely, messenger RNA in
prokaryotic cells can also be cut and rejoined. As the basic principles of such
a splicing reaction apply equally well to eukaryotic mRNA, the phenomenon is
described more fully in the sections which follow.
Related Topics