Chapter: Genetics and Molecular Biology: Protein Structure
Structures of DNA-Binding Proteins
Structures of DNA-Binding Proteins
A protein that regulates the expression of a gene
most often recognizes and binds to a specific DNA sequence in the vicinity of
that gene. Bacteria contain at least several thousand different genes, and most
are likely to be regulated. Eukaryotic cells contain at least 10,000 and
perhaps as many as 50,000 different regulated genes. Although combi-natorial tricks
could be used to reduce the number of regulatory proteins well below the total
number of genes, it seems likely that cells contain at least several thousand
different proteins that bind to specific se-quences. What must the structure of
a protein be in order that it bind with high specificity to one or a few
particular sequences of DNA? Does nature use more than one basic protein
structure for binding to DNA sequences?
We discussed the structure of DNA and pointed out
that the sequence of DNA can be read by hydrogen bonding to groups within the
major groove without melting the DNA’s double-stranded structure. Each of the
four base pair combinations, A-T, T-A, G-C, and C-G,
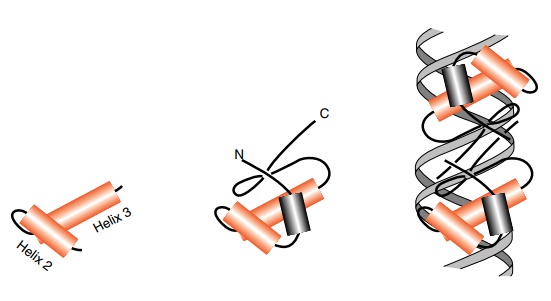
Figure
6.18 Left, a helix-turn-helix. Center,
the helix-turn-helix ofcrore-pressor
in the context of the remainder of the polypeptide chain. Right, a dimer of cro repressor with the recognition
helices fitting into the major groove of the DNA.
generates a unique pattern of hydrogen bond donors
and acceptors. Therefore, the identity of a base pair can be read by a single,
properly positioned, amino acid. Of course, an amino acid need not be
con-strained to bond only to a single base pair or just to the hydrogen bond
donors and acceptors in the major groove. Bonding from an amino acid residue is
also possible to multiple base pairs, to the deoxyribose rings, or the
phosphate groups.
Since the width of the major groove of DNA nicely
accepts an alpha-helix, it seems likely that such helices play a major role in
DNA-binding proteins, and this has been found. We might also expect proteins to
maximize their sequence selectivity by constructing their recognition surfaces
to be as rigid as possible. If the surface is held in the correct shape while the
protein is free in solution, then none of the protein-DNA binding energy needs
to be consumed in freezing the surface in the correct shape. All the
interaction energy between the protein and DNA can go into holding the protein
on the DNA and none needs to be spent holding the protein in the correct
conformation. Further, the DNA-contacting surface must protrude from the
protein in order to reach into the major groove of the DNA. These
considerations lead to the idea that proteins may utilize special mechanisms to
stiffen their DNA-binding domains, and indeed, this expectation is also met.
The high interest and importance in gene regulatory
proteins means that a number have been purified and carefully studied. Sequence
analysis and structure determination have revealed four main families of
DNA-binding domains. These are the helix-turn-helix, zinc domain, leucine
zipper, and the beta-ribbon proteins.
The helix-turn-helix domains possess a short loop
of four amino acids between the two helical regions (Fig. 6.18). Because the
connection between the helices is short and the helices partially lie across
one another, they form a rigid structure stabilized by hydrophobic interac-
tions between the helices. For historical reasons
the first of these two helices is called Helix 2, and the following helix is
called Helix 3. In these proteins Helix 3 lies within the major groove of the
DNA, but its orientation within the groove varies from one DNA-binding protein
to another. In some, the helix lies rather parallel to the major groove, but in
others, the helix sticks more end-on into the groove. The actual positioning of
the helix in the groove is determined by contacts between the protein and
phosphate groups, sugar groups, and more distal parts of the protein. Contacts
between the protein and DNA need not be direct. Trp repressor makes a substantial number of contacts indirectly
viawater molecules. Most prokaryotic helix-turn-helix proteins are
ho-modimeric, and they therefore bind a repeated sequence. Since the subunits
face one another, the repeats are inverted, and form a symmet-rical sequence,
for example AAAGGG-CCCTTT. Developmental genes in eukaryotes often are
regulated by proteins that are a close relative of the helix-turn-helix
protein. These homeodomain proteins possess struc-tures highly similar to the
helix-turn-helix structures, but either of the helices may be longer than their
prokaryotic analogs. The homeodo-main proteins usually are monomeric and
therefore bind to asymmetric sequences.

The zinc finger proteins are the most prominent
members of the zinc-containing proteins that bind to DNA. The zinc finger is a
domain of 25 to 30 amino acids consisting of a 12 residue helix and two
beta-ribbons packed against one another. In these proteins, Zn ligands to four
amino acids and forms a stiffening cross-bridge. Often the Zn is held by two
cysteines at the ends of the ribbons beside the turn, and by two histidines in
the alpha-helix. This unit fits into the major groove of the DNA and contacts
three base pairs. Usually these Zn-finger proteins
contain multiple fingers, and the next finger can
contact the following three base pairs. Other Zn-containing proteins that bind
to DNA contain multiple Zn ions in more complex structures. These proteins also
use residues in the alpha-helix and at its end to contact the DNA. Zn fingers
can also be used to make sequence-specific contacts to RNA.
The leucine zipper proteins are of a particularly
simple design. They contain two alpha-helical polypeptides dimerized by
hydrophobic faces on each. Characteristically, each polypeptide contains
leucine residues seven amino acids apart that form the dimerization faces.
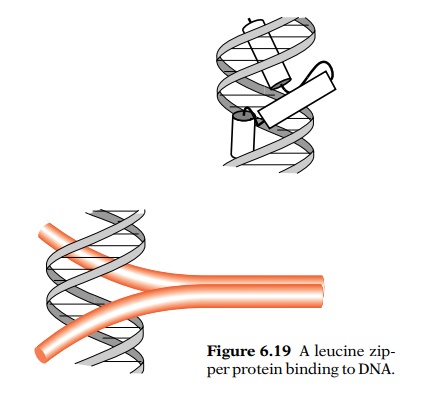
Beyond the dimerization domains the helices diverge as in a “Y”, with each arm
passing through a major groove (Fig. 6.19). Few leucine zipper proteins have
been identified in prokaryotes, but they are common in eukaryotes. They are
notable in forming heterodimers. For example, the fos-jun transcription factor is a leucine zipper protein. It is
easy to see how specific dimerization could be determined by patterns of
positive and negative charges near the contact regions of the two dimerization
regions.
The beta-sheet domains contact either the major or
minor groove of DNA via two antiparallel beta-strands. A few prokaryotic and
eukaryotic examples of these proteins are known, the most notable being the
X-ray determined structure of the MetJ repressor-operator complex. At least
part of the DNA contacts made by transcription factor TFIID are in the minor
groove and are made by such beta-strands.
Related Topics