Chapter: Plant Biochemistry: Multiple signals regulate the growth and development of plant organs
Light sensors regulate growth and development of plants
Light sensors regulate growth and development of plants
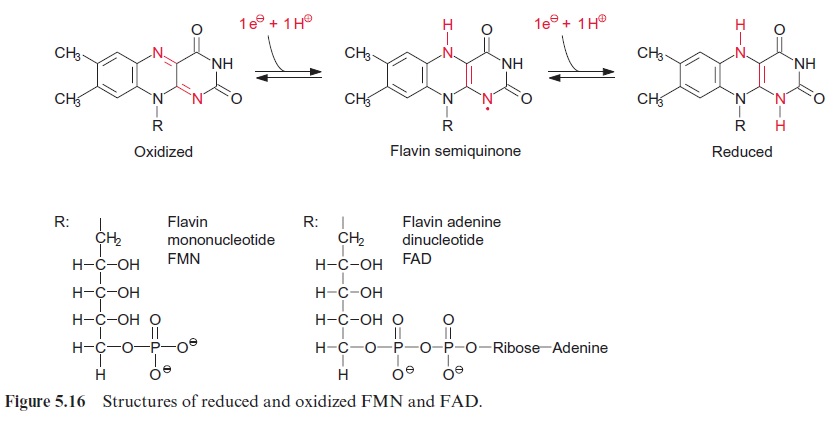
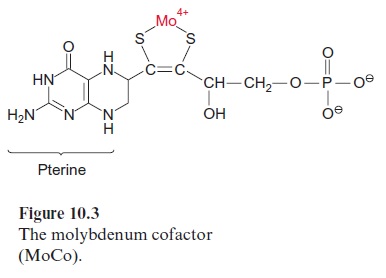
Light controls plant development from germination to the formation of flowers in many different ways. Important light sensors are thephytochromes which sense red light. Phytochromes are involved when light initiates the germination and greening of the seedling and in the adaptation of the pho-tosynthetic apparatus of the leaves to full sunlight or shade. Five different phytochromes (A–E) have been identified in the model plant Arabidopsis thaliana . Plants also have photoreceptors for blue and UV light for their adaptation to the full spectrum of sunlight. So far, three proteins have been identified as blue light receptors; these are cryptochromes 1 and 2, each comprising a flavin (Fig. 5.16) and a pterin (Fig. 10.3); and phototropin, containing one flavin as a blue light-absorbing pigment.
Phytochromes function as sensors for red light
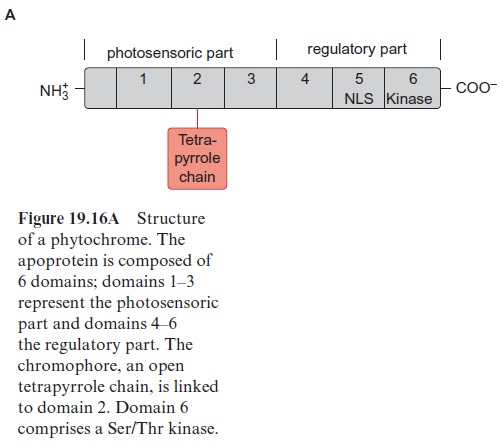
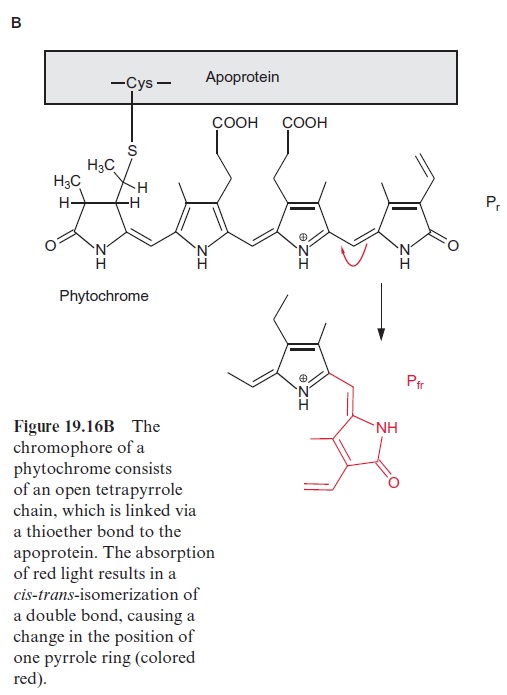
Since the structure and function of phytochromes have been studied exten-sively in the past, they offer a good example for a detailed discussion on the problems of signal transduction in plants. Phytochromes are soluble dimeric proteins. The monomer consists of an apoprotein (molecular mass 120–130 kDa) with six domains (Fig. 19.16A). The first three domains repre-sent the photosensoric part of the protein. These domains are also present in bacterial and fungal phytochromes. The remaining domains 4, 5, and 6 form the regulatory part. Plant phytochromes contain a sequence at the N-terminus of the apoprotein, which is involved in the Pfr → Pr conversion. Domain 6 possesses a kinase which binds ATP and catalyzes an autophosphorylation of the phytochrome. Domain 2 contains an open chain tetrapyrrole that is linked to the protein via a sulfhydryl group of a cysteine residue. It is the chromophore of the holoprotein (Fig. 19.16B). The autocatalytic binding of the tetrapyrrole to the apoprotein results in the formation of a phytochrome Pr (r=red) with an absorption maximum at about 660 nm (red light) (Fig. 19.17). The absorption of this light results in a change in the chromophore; a double bond between the two pyrrole rings changes from the trans- to the cis-configuration (colored red in Fig. 19.16B), which subsequently changes the conformation of the protein. The phytochrome in this new conformation has an absorption maximum at about 730 nm (far-red light) named Pfr, rep-resenting the active form of the phytochrome. It reflects the state of illumina-tion. Pfr is reconverted to Pr by the absorption of far-red light. Since the light absorption of Pr and Pfr overlaps (Fig. 19.17), depending on the color of the irradiated light, a reversible equilibrium between Pr and Pfr is attained. Thus, with light of 660 nm, 88% of the total phytochromes are present as Pfr and at 720 nm, only 3% is in the Pfr form. In bright sunlight, where the red com-ponent is stronger than the far-red component, the phytochrome is present primarily as Pfr and indicates the state of illumination to the plant.
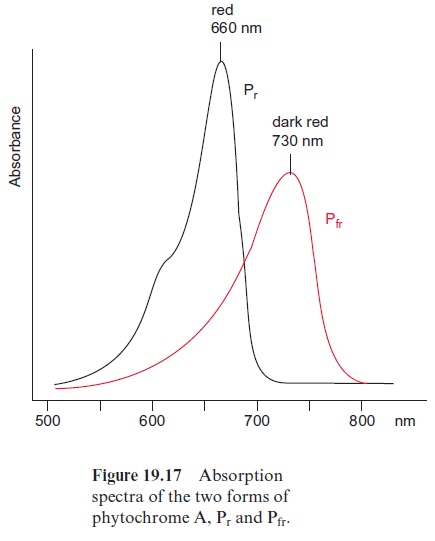
Whereas the inactive form of phytochrome (Pr) has quite a long lifetime (~100 h), the active form (Pfr) is converted within 30 to 60 minutes. Pfr can be recovered by a reversal of its formation (Fig. 19.18). In the case of phy-tochrome A, however, the light absorption can be terminated by conjuga-tion of Pfr with ubiquitin, which marks it for proteolytic degradation by the proteasome pathway.
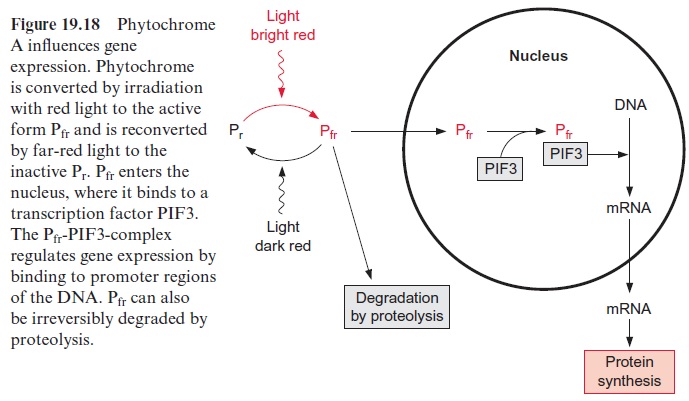
Phytochromes undergo a photoconversion upon illumination with red and far-red light. In addition to this, phytochromes are transferred between the cytosol and the nucleus. The C-terminal domain 5 (Fig. 19.16A) contains a nuclear localization signal(NLS) that is responsible for targeting to the nucleus. Under the influence of light the protein kinase of domain 6 causes an autophosphorylation of the protein. This phosphorylation releases the phytochrome from a cytosolic anchor and the NLS domain is uncovered. This allows the migration of the Pfr form of phytochrome into the nucleus, where it associates with factors, such as thephytochrome interacting factor (PIF3) and other transcription factors to affect gene expression (Fig. 19.18). It has been shown that phytochrome A affects the transcription of 10% of all genes in Arabidopsis. This effect of phytochromes is modulated by phytohormones, such as cytokinins and brassinosteroids, and the signal cascades of pathogen defense. Light sensors, phytohormones, and defense reactions appear to be interwoven in a cellular network.
Phytochrome A, having been most thoroughly investigated, has an absorption maximum at far-red light and is instable in light, whereas the other phytochromes (B–E) are light stable. Phytochrome A often forms homodimers, while the other phytochromes aggregate to heterodimers. The functions of phytochromes B–E have still to be resolved in detail.
Phototropin and cryptochromes are blue light receptors
Light sensors such as phototropin and cryptochrome are present in plant cells in order to use the blue spectrum (320–500 nm) of the sun efficiently. Blue light induces stoma opening, the elongation of the hypocotyls and the chloroplast movement. At low blue light intensity the chloroplasts move to that part of the cell exposed to the light in order to optimize light harvest-ing, whereas at higher blue light intensity the chloroplasts retreat from the light source to avoid destruction by excessive light.
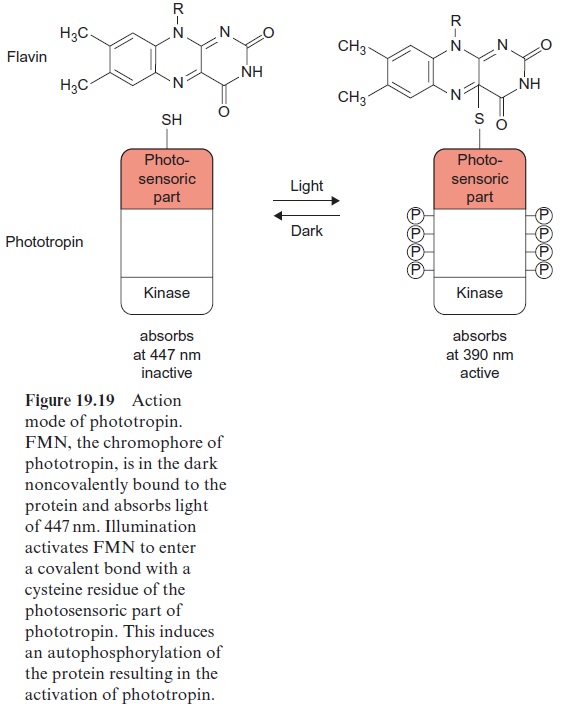
Phototropin was first isolated in 1997 from pea cotyledons that had been irradiated with blue light, and was found to be ubiquitous in higher plants. The protein structure of phototropin shows similarities to the structure of phytochromes. As in phytochromes the photosensor is located at the N-terminal region which contains a flavine mononucleotide (FMN) (Fig. 5.16) as light absorbent. Moreover, there is a serine/threonine kinase at the C-terminal region, catalyzing the autophosphorylation of the pro-tein. In the dark FMN is associated noncovalently (Fig. 19.19). Upon illu-mination the FMN is covalently bound within microseconds to a cysteine residue that changes the absorption maximum of the flavine from 447 nm to 390 nm. This induces the kinase at the C-terminal region to catalyze the autophosphorylation of 8 serine residues, converting the phototropin into an active state. In the dark the FMN is released again, the phosphate groups are split off by hydrolysis, presumably by a phosphatase, and the phototropin returns within 10–100 s to the inactive ground state.
Cryptochromes also absorb blue light but are not structurally related to phototropin. They derive from microbial DNA repair enzymes, so-called photolyases. They are flavoproteins influencing many processes in plants, such as hypocotyl and stem elongation, regulation of flowering time and anthocyan accumulation. Details of the interplay of the two blue light receptors, phototropins and cryptochromes, in plant development and metabolism are still largely unknown.
Related Topics