Chapter: Biology of Disease: Genetic Diseases
Detecting, Diagnosing and Screening Human Genetics Diseases
DETECTING, DIAGNOSING AND SCREENING HUMAN GENETIC DISEASES
Identifying the genetic basis for a human disorder usually requires an analysis of the family history as far back as possible. If this shows that the trait is inherited, it is possible to predict whether the mutant allele is dominant or recessive and whether it is X- or autosomal-linked. Obviously, dominant mutations are the simplest to detect. If they are X-linked, then affected fathers will pass the trait to all their daughters. However, if autosomal, then dominant mutations would be expected to occur in approximately 50% of the children of an affected heterozygous individual.
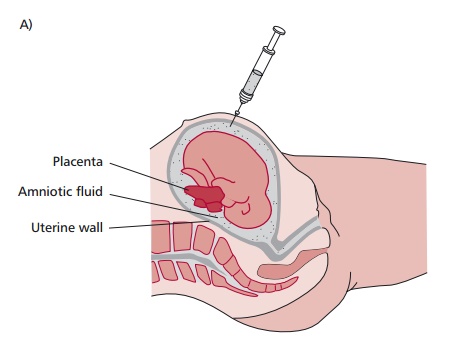
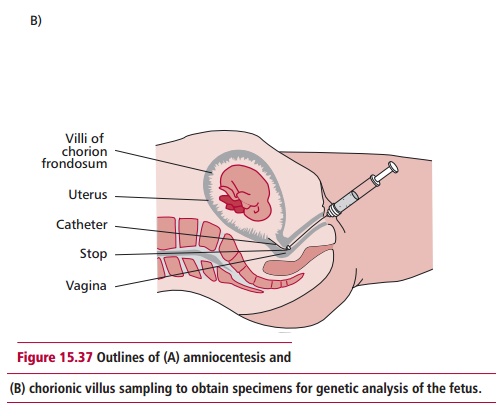
In many cases, the diagnosis of a genetic disease can be made prenatally. The most widely used methods for this are amniocentesis and chorionic villus sampling (CVS). In the former, a needle is used to withdraw amniotic fluid (Figure 15.37 (A)) containing cells genetically identical to those of the fetus. In CVS, a catheter is inserted into the uterus and a small tissue sample of the fetal chorion removed (Figure 15.37 (B)). In both cases, a variety of cytogenetic and biochemical tests can then be performed on the cells and tissue to identify any single gene disorders or chromosomal abnormalities.
People in whom a genetic defect has been detected will be advised to seek genetic counseling, especially if they wish to have children. The advice they are given is based on analyses of the risks that they may produce a child with a genetic abnormality or that they themselves may develop a late onset genetic disease. A broad range of information is required and genetic counselors require training to provide this information with appropriate consideration.
DETECTION OF HEREDITARY DISEASES
Tests based on recombinant DNA technology have increased in recent years because of their high sensitivity and accuracy in detecting genetic disorders at the prenatal stage. The detection of hereditary diseases in a given genome was traditionally a long and laborious task. However, modern techniques of chromosome and DNA analyses have resulted in a dramatic increase in the number of tests for genetic diseases with a considerable improvement in the time required for analysis. For example, using PCR, polymerase chain reaction , virtually any portion of a gene or even whole genes can be amplified for analysis by electrophoretic techniques or sequenced to detect mutations. This not only allows for diagnosing patients with inherited disorders but also the detection of mutations in carriers, even though they do not express any symptom of the disorder. It is also a direct way of distinguishing different mutations within a single gene, each of which can lead to disorders, for example muscular dystrophies which are described. It is hoped that eventually predictive tests for disorders that have only some genetic component, such as heart disease and cancers, will eventually be developed. Indeed, PCR analysis of cells lost in feces has demonstrated premalignant changes in the gastrointestinal tract and allowed patients at risk of developing colon cancer to be identified. This is of clinical importance since the earlier treatments of malignant conditions are started, the more favorable the prognosis.
Cloned DNA sequences have expanded the range of prenatal testing because they allow the fetal genotype to be examined directly, rather than relying on secondary tests for the products of the normal or mutant genes. Thus, mutations in DNA can be detected in those cases where an aberrant product cannot be detected prior to birth, even though a test is available. A test for sickle cell anemia using a restriction enzyme has been described. However, RFLP analysis requires that the mutation in the gene alters therestriction sites recognized by restriction endonucleases and this is not always the case. In contrast, allele-specific oligonucleotides (ASOs) are synthetic nucleotide probes, which bind only to their complementary DNA. They will not hybridize to other sequences and, in appropriate highly stringent conditions, differentially bind to and distinguish between alleles that differ by as little as one nucleotide. They can thus distinguish between the native and mutant forms of a gene with excellent resolution and potential versatility.
Tests based on ASOs are now available to screen for mutations associated with cystic fibrosis and glaucoma.
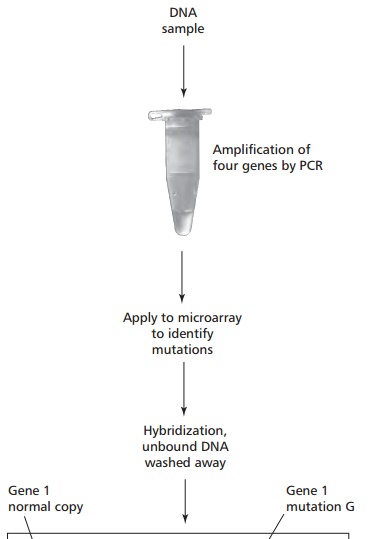
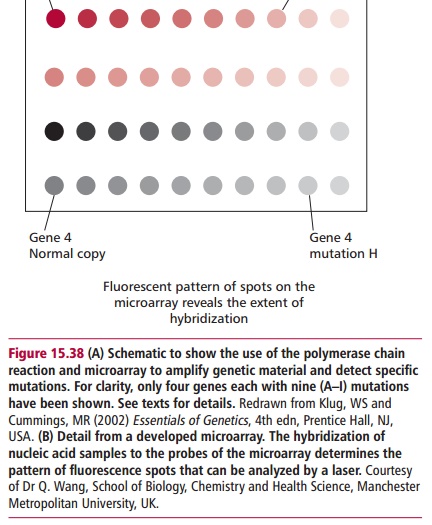
Allele-specific oligonucleotides are being increasingly combined with DNA microarrays, or chips. These are small glass slides or nylon membranes divided into squares, called fields, each of which contains a specific probe about 20 oligonucleotides long bound to the slide. DNA microarrays can be produced containing thousands of probes and can thus be used to analyze many different genes simultaneously. DNA is extracted from the cells and digested with one or more restriction endonucleases to produce small sized fragments that are labeled with a fluorescent dye. These are heated to separate the DNA into single strands which are then added to the microarray. Fragments whose sequence is complementary to a probe will hybridize with it and bind to the microarray; others will be washed off even if their complementary sequence differs from the probe by as little as one base. A laser based device is used to scan the array and determine where the binding has occurred to identify specific mutations (Figure 15.38). DNA chips are in development that will contain probes for all of the approximately 22 000 human genes to allow the simultaneous analysis of the DNA from a single person for hundreds of genetic diseases or genetic predispositions. The generation of such data has profound social, ethical and legal implications.
Potential parents in whom a genetic defect is detected must be advised to have genetic counseling. The advice they are given is based on analyses of the risks of them having a child with a genetic abnormality or that they may develop a late onset genetic disease. Information drawn from a wide range of sources is required in such circumstances and genetic counselors must provide such information with tact and sensitivity.
Related Topics