Chapter: Biotechnology Applying the Genetic Revolution: Recombinant DNA Technology
Specific Types of Cloning Vectors
SPECIFIC
TYPES OF CLONING VECTORS
Because E. coli is the main host organism used for manipulating DNA, most
vectors are based on plasmids or viruses that can survive in E. coli or similar bacteria. Most
vectors have bacterial origins of replication and antibiotic resistance genes.
The polylinker or multiple cloning site is usually placed between prokaryotic
promoter and terminator sequences. Some vectors may also supply the ribosome
binding site, so that any inserted coding sequence will be expressed as a
protein. Many other features are present in specialized cloning vectors.
The following discussion will
introduce some of the different categories of vectors with their essential
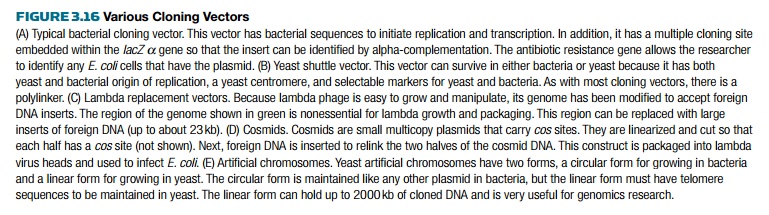
features (Fig. 3.16).
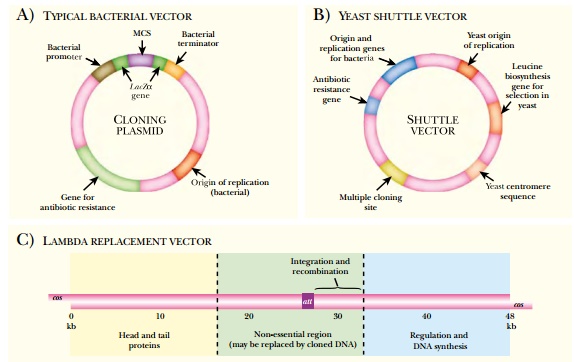
Many vectors are based on the
2µ circle of yeast. The native version of the 2µ circle has been modified in a variety of ways for use as
a cloning vector. A shuttle vector contains origins of replication for two
organisms plus any other sequences necessary to survive in either organism (see Fig.
3.16B). Shuttle vectors that are based on the 2µ plasmid have the components needed
for survival in yeast and bacteria, plus antibiotic resistance and a
polylinker. The Cen sequence is a eukaryotic centromere (Cen) sequence that keeps the
plasmid in the correct location during mitosis and meiosis in yeast. Because
yeast cells are eukaryotic and also have such a thick cell wall, most
antibiotics do not kill yeast. Therefore, a different strategy is used to
detect the presence of plasmids in yeast. A gene for synthesis of an amino
acid, such as leucine, allows strains of yeast that require leucine to grow.
Bacteriophage vectors are viral genomes that have been modified so
that large pieces of nonviral DNA
can be packaged in the virus particle. Lambda bacteriophages have linear
genomes with two cohesive ends—cos sequences (lambda cohesive ends). These are 12-base overlapping
sticky ends. When inside the virus coat, the cohesive ends are coated with
protein to prevent them from annealing. After lambda attaches to E. coli, it inserts just the linear DNA.
The proteins that protect the cohesive ends are lost, and the genome
circularizes with the help of DNA ligase. The circular form is the replicative form (RF), and it
replicates by the rolling circle mechanism. Expression of various lambda genes
produces the proteins that assemble into protein coats. Each coat is packaged
with one genome, and after many of these are assembled, the E. coli host explodes, releasing the new
bacteriophage to infect other cells.
The lambda bacteriophage is a
widely used cloning vector (see Fig. 3.16C). The middle segment of the lambda
genome has been deleted and a polylinker has been added.
An insert of 37 to 52 kb can
be ligated into the polylinker and packaged into viral particles. In order to
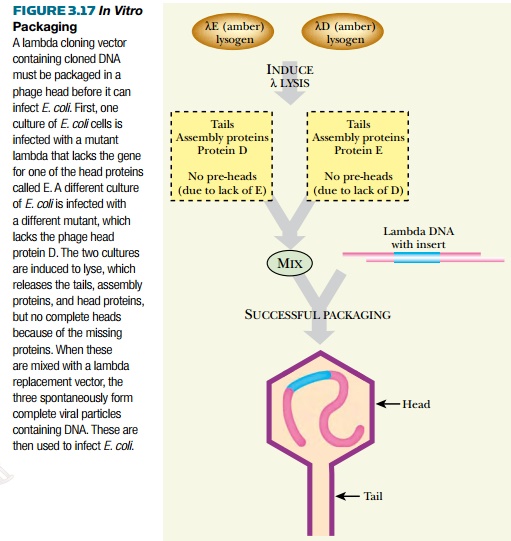
work with the bacteriophage DNA without killing the entire E. coli culture, one or more genes necessary for packaging are
deleted. When the researcher wants to form fully packaged bacteriophages, coat
proteins from helper virus can be
added
(Fig. 3.17). The helper
viruses do not contain foreign DNA, but supply the missing genes for the coat
proteins. Because coat proteins self-assemble in vitro, helper lysates are mixed with recombinant lambda DNA and
complete virus particles containing DNA are produced. This is known as in vitro packaging.
Cosmid vectors can hold pieces of DNA up to 45 kb in length (see Fig. 3.16D). These are highly modified lambda vectors with all the sequences between the cos sites removed and replaced with the insert. The DNA of interest is ligated between the two cos sites using restriction enzymes and ligase. This construct is packaged into a lambda particle produced by helper phage as shown earlier (see Fig. 3.17), and then these are used to infect E. coli.
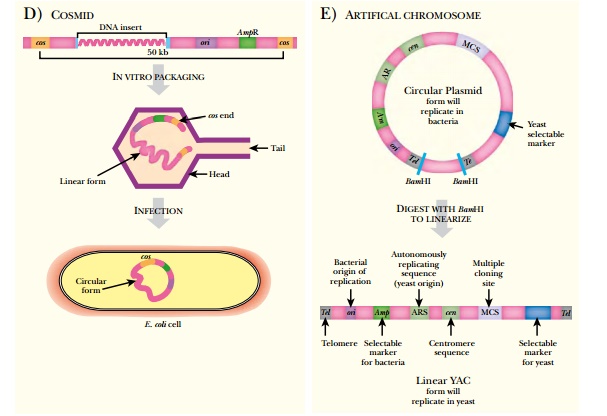

Artificial chromosomes hold the largest pieces of DNA (see Fig.
3.16E). These include yeast
artificial chromosomes (YACs), bacterial artificial chromosomes (BACs), and P1
bacteriophage artificial chromosomes (PACs). They are used to contain lengths
of DNA from 150 kb to 2000 kb. YACs hold the largest amount of DNA, up to about
2000 kb. YACs have yeast centromeres and yeast telomeres for maintenance in
yeast. BACs can be circularized and grown in bacteria; therefore, they have a
bacterial origin of replication and antibiotic resistance genes. The
flexibility of artificial chromosomes makes them most useful for sequencing
entire genomes, especially those of higher organisms with vast amounts of DNA.
Related Topics