Chapter: Biotechnology Applying the Genetic Revolution: Recombinant DNA Technology
Constructing a Library of Genes
CONSTRUCTING
A LIBRARY OF GENES
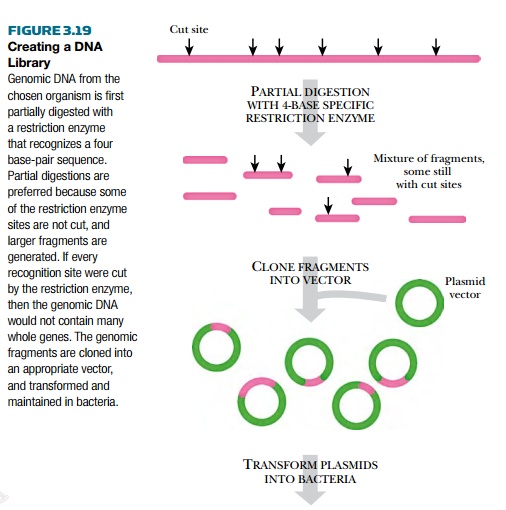
Gene libraries are used to
find new genes, to sequence entire genomes, and to compare genes from different
organisms (Fig. 3.19). Gene libraries are made when the entire DNA from one
particular organism is digested into fragments using restriction enzymes, and
then each of the fragments is cloned into a vector and transformed into an appropriate
host.
The basic steps used to
construct a library are:
1.
Isolate the chromosomal DNA from an organism, such as E. coli, yeast, or humans.
2.
Digest the DNA with one or two different restriction enzymes.
3.
Linearize a suitable cloning vector with the same restriction
enzyme(s).
4.
Mix the cut chromosome fragments with the linearized vector and
ligate.
5.
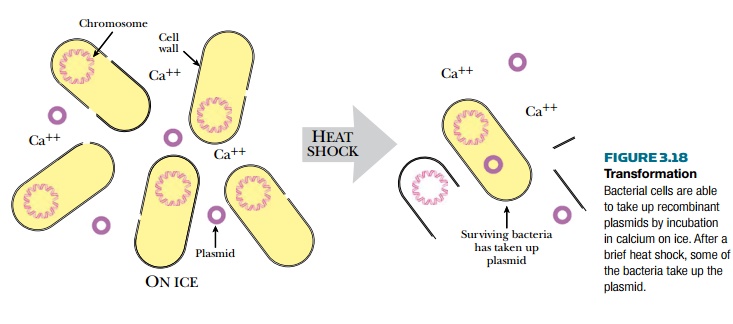
Transform this mixture into E.
coli.
6.
Isolate large numbers of E.
coli transformants.
The type of restriction enzyme affects the type of library. Because restriction sites are not evenly spaced in the genome, some inserts will be large and others small. Using a restriction enzyme that recognizes only four base pairs will give a mixture of mostly small fragments, whereas a restriction enzyme that has a six or eight base-pair recognition sequence will generate larger fragments. (Note that finding a particular four base-pair sequence in a genome is more likely than finding a six base-pair sequence.) Even if an enzyme that recognizes a four base-pair recognition sequence is used to digest the entire genome, there may still be segments that are too large to be cloned. Conversely, clustered restriction sites will cause some genes will be cut into several pieces. To avoid this, partial digestion is often used. The enzyme is allowed to cut the DNA for only a short time, and many of the restriction enzyme sites are not cut, leaving larger pieces for the library. In addition, it is usual to construct another library using a different restriction enzyme.
Related Topics