Chapter: Biotechnology Applying the Genetic Revolution: Recombinant DNA Technology
Fluorescence in Situ Hybridization (FISH)
FLUORESCENCE
IN SITU HYBRIDIZATION (FISH)
The previously discussed
hybridization techniques rely on purified DNA or RNA run in an agarose gel. In fluorescence in situ hybridization (FISH), the probe is
hybridized directly to DNA or RNA within the cell. As described earlier, the
probe is a small segment of DNA that has been labeled with fluorescent tags in
order to be visualized. The target DNA or RNA is located within the cell and
requires some special processing. The target cells may be extremely thin
sections of tissue from a particular organism. For example, when a person has a
biopsy, a small piece of tissue is removed for analysis. This tissue is
preserved and then cut into extremely thin sections to be analyzed under a
microscope. These can be used to determine the presence of a gene with FISH.
Another source of target cells for FISH is cultured mammalian or insect cells.
Additionally, blood can be isolated and processed to isolate the white blood
cells. (Note: Red blood cells do not contain a nucleus and therefore do not
contain DNA.) Chromosomes from white blood cells can be isolated and dropped
onto a glass slide. Blood sample DNA may be found both in interphase, with the
chromosomes spread out, or in metaphase, with the chromosomes highly condensed.
Whether the target DNA is
from blood, cells cultured in dishes, or actual tissue sections, the cells must
be heated to make the DNA single-stranded. Samples where RNA is the target do
not need to be heated. The fluorescently labeled probe hybridizes to
complementary sequences in the DNA or RNA, and when the cells are illuminated
at the appropriate wavelength, the probe location on the chromosome can be
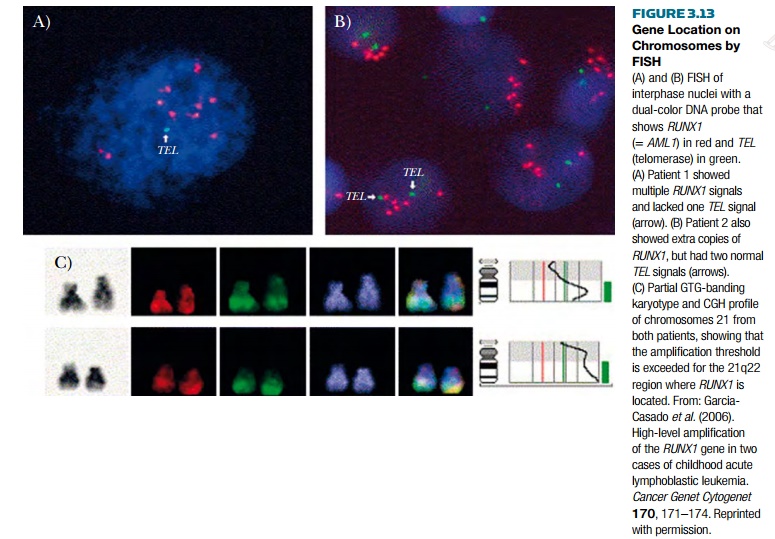
identified by fluorescence. Figure 3.13 shows FISH analysis of the RUNX1 gene that is amplified in certain
cases of human acute leukemia due to polysomy (that is, multiple copies) of
chromosome 21.
Related Topics