Chapter: Biotechnology Applying the Genetic Revolution: DNA Synthesis in Vivo and in Vitro
Randomly Amplified Polymorphic DNA
RANDOMLY
AMPLIFIED POLYMORPHIC DNA
Another PCR application, called randomly amplified polymorphic DNA (RAPD), allows the researcher to compare the genetic relatedness of two DNA samples (Fig. 4.25). First, two DNA samples are isolated from two different organisms. Thus there are two samples of target DNA, which are compared by using the same set of primers. As before, the sequence information is unknown, but rather than using degenerate primers, primers with a randomly chosen sequence are made. The length of a primer determines how often it will bind within the target DNA. If a particular primer were, say, 5 bases long, it would bind once on average every 45 bases = 4 × 4 × 4 × 4 × 4 = 1024 bases. If the target DNA were a sample from a large genome, such a primer would bind far too many times. In practice longer primers of around 10 bases are often suitable. The random primer is mixed with nucleotides, Taq polymerase, and each of the target DNA samples as for normal PCR reactions. In order to amplify any target DNA fragments, two of the random primers must bind to the target DNA, on opposite strands, usually within a few thousand bases. The results of the two PCR reactions are compared using gel electrophoresis. The number and size of PCR products will vary for the two samples. If two organisms are very closely related, then their DNA will be close in sequence. Hence, the PCR products will be very
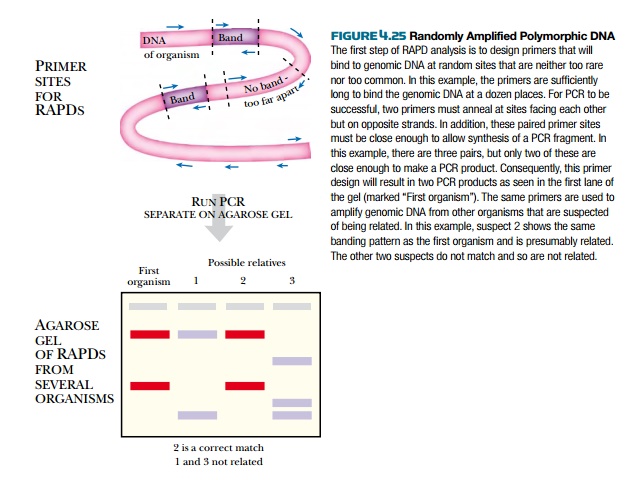
similar with only one or two
different fragments. If the pattern of PCR products is totally different, then
the two organisms are not related. Comparing RAPDs from two organisms can thus
give an estimate of relatedness.
Related Topics