Molecular Genetics - Packaging of DNA helix | 12th Zoology : Chapter 5 : Molecular Genetics
Chapter: 12th Zoology : Chapter 5 : Molecular Genetics
Packaging of DNA helix
Packaging
of DNA helix
The distance between two
consecutive base pairs is 0.34nm (0.34×10-9m) of the DNA double
helix in a typical mammalian cell. When the total number of base pairs is
multiplied with the distance between two consecutive base pairs (6.6 × 10-9 ×
0.34 ×10-9 m/bp), the length of DNA double helix is
(The total length of the double helical DNA = total number of base pairs ×
distance between two consecutive base pairs). If the length of E. coli
DNA is 1. 36 mm, the number of base pairs in E. coli is 4 ×106m
(1.36 × 103 m/0.34 ×10-9 ). The length of the DNA double
helix is far greater than the dimension of a typical mammalian nucleus
(approximately 10-6 m) . How is such a long DNA polymer packaged in
a cell?
Chromosomes are carriers
of genes which are responsible for various characters from generation to
generation. Du Praw (1965) proposed a single stranded model (unineme), as a
long coiled molecule which is associated with histone proteins in eukaryotes.
Plants and animals have more DNA than bacteria and must fold this DNA to fit
into the cell nucleus. In prokaryotes such as E. coli though they do not
have defined nucleus, the DNA is not scattered throughout the cell. DNA (being
negatively charged) is held with some proteins (that have positive charges) in
a region called the nucleoid. The DNA as a nucleoid is organized into large
loops held by protein. DNA of prokaryotes is almost circular and lacks
chromatin organization, hence termed genophore.
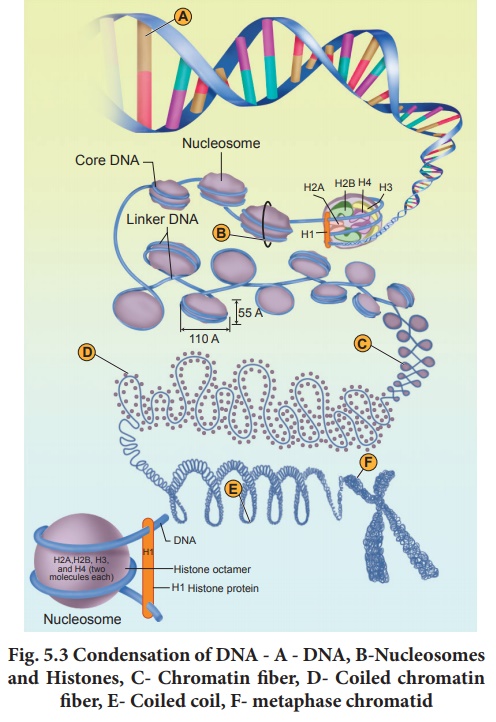
In eukaryotes, this
organization is much more complex. Chromatin is formed by a series of repeating
units called nucleosomes. Kornberg proposed a model for the nucleosome,
in which 2 molecules of the four histone proteins H2A, H2B, H3 and H4 are
organized to form a unit of eight molecules called histone octamere.
The negatively charged DNA is wrapped around the positively charged
histone octamere to form a structure called nucleosome. A typical
nucleosome contains 200 bp of DNA helix. The histone octameres are in
close contact and DNA is coiled on the outside of nucleosome. Neighbouring
nucleosomes are connected by linker DNA (H1) that is exposed to enzymes. The
DNA makes two complete turns around the histone octameres and the two turns are
sealed off by an H1 molecule. Chromatin lacking H1 has a beads-on-a-string appearance
in which DNA enters and leaves the nucleosomes at random places. H1 of
one nucleosome can interact with H1 of the neighbouring nucleosomes resulting
in the further folding of the fibre. The chromatin fiber in interphase nuclei
and mitotic chromosomes have a diameter that vary between 200 -300 nm and
represents inactive chromatin. 30 nm fibre arises from the folding of
nucleosome, chains into a solenoid structure having six nucleosomes per
turn. This structure is stabilized by interaction between different H1
molecules. DNA is a solenoid and packed about 40 folds. The hierarchical nature
of chromosome structure is illustrated in (Fig. 5.3). Additional
set of proteins are required for packing of chromatin at higher level
and are referred to as non-histone chromosomal proteins (NHC). In a typical
nucleus, some regions of chromatin are loosely packed (lightly stained) and are
referred to as euchromatin. The chromatin that is tightly packed (stained
darkly) is called heterochromatin. Euchromatin is transcriptionally active and
heterochromatin is transcriptionally inactive.
![]()
![]()
Related Topics