Steps, Application - Molecular Genetics - DNA fingerprinting technique | 12th Zoology : Chapter 5 : Molecular Genetics
Chapter: 12th Zoology : Chapter 5 : Molecular Genetics
DNA fingerprinting technique
DNA
fingerprinting technique
The DNA fingerprinting
technique was first developed by Alec Jeffreys in 1985 (Recipient of the Royal
Society’s Copley Medal in 2014). Each of us have the same chemical structure of
DNA. But there are millions of differences in the DNA sequence of base pairs.
This makes the uniqueness among us so that each of us except identical twins is
different from each other genetically. The DNA of a person and finger prints
are unique. There are 23 pairs of human chromosomes with 1.5 million pairs of
genes. It is a well known fact that genes are segments of DNA which differ in
the sequence of their nucleotides. Not all segments of DNA code for proteins,
some DNA segments have a regulatory function, while others are intervening sequences
(introns) and still others are repeated DNA sequences. In DNA fingerprinting,
short repetitive nucleotide sequences are specific for a person. These
nucleotide sequences are called as variable number tandem repeats
(VNTR).The VNTRs of two persons generally show variations and are
useful as genetic markers.
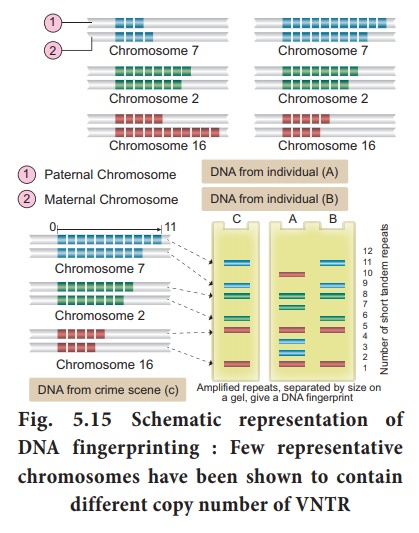
DNA finger printing
involves identifying differences in some specific regions in DNA sequence
called repetitive DNA, because in these sequences, a small stretch of
DNA is repeated many times. These repetitive DNA are separated from bulk
genomic DNA as different peaks during density gradient centrifugation. The bulk
DNA forms a major peak and the other small peaks are referred to as satellite
DNA. Depending on base composition (A : T rich or G : C rich), length of
segment and number of repetitive units, the satellite DNA is classified into
many sub categories
such as micro-satellites, mini-satellites, etc., These sequences do not code
for any proteins, but they form a large portion of human genome. These
sequences show high degree of polymorphism and form the basis of DNA
fingerprinting (Fig. 5.15). DNA isolated from blood, hair, skin cells,
or other genetic evidences left at the scene of a crime can be compared through
VNTR patterns, with the DNA of a criminal suspect to determine guilt or
innocence. VNTR patterns are also useful in establishing the identity of a
homicide victim, either from DNA found as evidence or from the body itself.
The Steps in DNA
Fingerprinting technique is depicted in Fig. 5.16.
1.
Extraction of DNA
The process of DNA
fingerprinting starts with obtaining a sample of DNA from blood, semen, vaginal
fluids, hair roots, teeth, bones, etc.,
2.
Polymerase chain reaction (PCR)
In many situations,
there is only a small amount of DNA available for DNA fingerprinting. If needed
many copies of the DNA can be produced by PCR (DNA amplification).
3.
Fragmenting DNA
DNA is treated with
restriction enzymes which cut the DNA into smaller fragments at specific sites.
4.
Separation of DNA by electrophoresis
During electrophoresis in an agarose gel, the DNA fragments are
separated into bands of different sizes. The bands of separated DNA are sieved
out of the gel using a nylon membrane (treated with chemicals that allow for it
to break the hydrogen bonds of DNA so there are single strands).
5.
Denaturing DNA
The DNA on gels is
denatured by using alkaline chemicals or by heating.
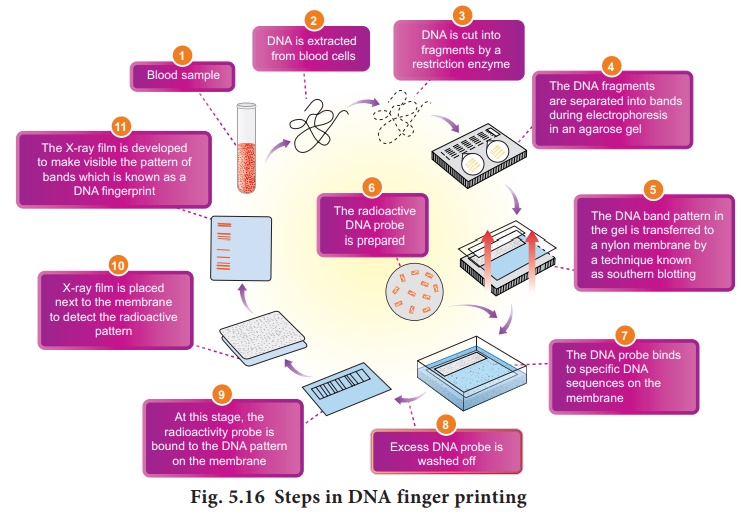
6.
Blotting
The DNA band pattern in
the gel is transferred to a thin nylon membrane placed over the ‘size
fractionated DNA strand’ by Southern blotting.
7.
Using probes to identify specific DNA
A radioactive probe (DNA labeled with a radioactive substance) is
added to the DNA bands. The probe attaches by base pairing to those restriction
fragments that are complementary to its sequence. The probes can also be
prepared by using either ‘fluorescent substance’ or ‘radioactive isotopes’.
8.
Hybridization with probe
After the probe
hybridizes and the excess probe washed off, a photographic film is placed on
the membrane containing ‘DNA hybrids’.
9.
Exposure on film to make a genetic/ DNA Fingerprint
The radioactive label
exposes the film to form an image (image of bands)corresponding to specific DNA
bands. The thick and thin dark bands form a pattern of bars which
constitutes a genetic fingerprint.
Application of DNA finger printing
·
Forensic analysis - It can be used in the identification of
a person involved in criminal activities, for settling paternity or maternity
disputes, and in determining relationships for immigration purposes.
·
Pedigree analysis – inheritance pattern of genes through
generations and for detecting inherited diseases.
·
Conservation of wild life – protection of endangered species. By
maintaining DNA records for identification of tissues of the dead endangered
organisms.
·
Anthropological studies–It is useful in determining the origin
and migration of human populations and genetic diversities.
Related Topics