Molecular Genetics - DNA Replication | 12th Zoology : Chapter 5 : Molecular Genetics
Chapter: 12th Zoology : Chapter 5 : Molecular Genetics
DNA Replication
DNA
Replication
Replication of DNA takes
place during the S phase of cell cycle. During replication, each DNA molecule
gives rise to two DNA strands, identical to each other as well as to the parent
strand. Three hypotheses of DNA replication have been proposed. They are
conservative replication, dispersive replication, and semi-conservative
replication.
In conservative replication, the original double helix serves as a template. The original molecule is preserved intact and an entirely new double stranded molecule is synthesized. In dispersive replication, the original molecule is broken into fragments and each fragment serves as a template for the synthesis of complementary fragments. Finally two new molecules are formed which consist of both old and new fragments.
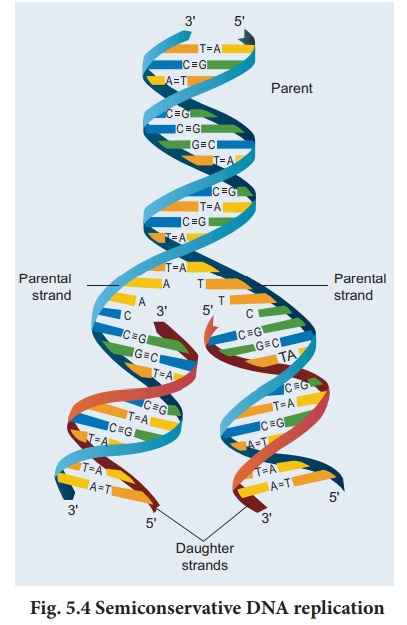
Semi-conservative
replication was proposed by Watson and Crick in 1953. This mechanism of
replication is based on the DNA model. They suggested that the two
polynucleotide strands of DNA molecule unwind and start separating at one end.
During this process, covalent hydrogen bonds are broken. The separated single
strand then acts as template for the synthesis of a new strand. Subsequently,
each daughter double helix carries one polynucleotide strand from the parent
molecule that acts as a template and the other strand is newly synthesised and
complementary to the parent strand (Fig. 5.4).
1. Experimental proof of DNA replication
The mode of DNA
replication was determined in 1958 by Meselson and Stahl. They designed an
experiment to distinguish between semi conservative, conservative and
dispersive replications. In their experiment, they grew two cultures of E.coli
for many generations in separate media. The ‘heavy’ culture was grown in a
medium in which the nitrogen source (NH4 Cl) contained the heavy
isotope 15N and the ‘light’ culture was grown in a medium in which the nitrogen
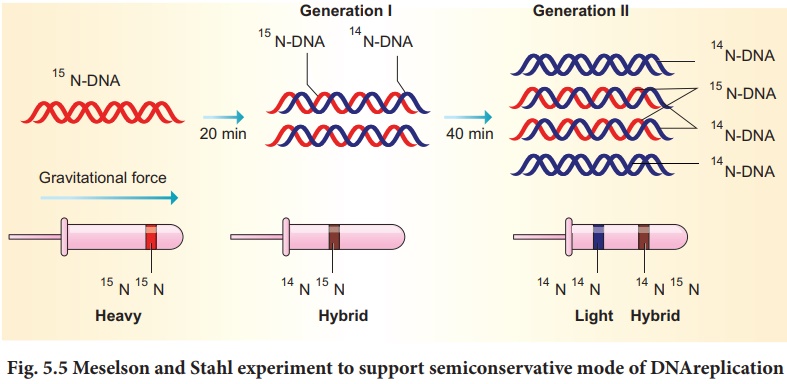
At the end of growth, they observed
that the bacterial DNA in the heavy culture contained only 15N and
in the light culture only 14N. The heavy DNA could be distinguished
from light DNA (15N from 14N) with a technique called Cesium
Chloride (CsCl) density gradient centrifugation. In this process,
heavy and light DNA extracted from cells in the two cultures settled into two
distinct and separate bands (hybrid DNA) (Fig. 5.5).
The heavy culture (15N)
was then transferred into a medium that had only NH 4Cl, and took
samples at various definite time intervals (20 minutes duration). After the
first replication, they extracted DNA and subjected it to density gradient
centrifugation. The DNA settled into a band that was intermediate in position
between the previously determined heavy and light bands. After the second
replication (40 minutes duration), they again extracted DNA samples, and this
time found the DNA settling into two bands, one at the light band position and
one at intermediate position. These results confirm Watson and Crick’s semi conservative
replication hypothesis.
2.
Enzymes and mechanism of replication
In prokaryotes,
replication process requires three types of DNA polymerases (DNA polymerase I,
II, and III). DNA polymerase is the main enzyme involved in DNA replication.
DNA polymerase I (also known as Kornberg enzyme) and DNA polymerase II
are involved in DNA repair mechanism. Eukaryotes have five types of DNA
polymerases that catalyses the polymerization of nucleotides at the 3' OH of
the new strand within a short period of time. E.coli that has 4.6 X 106
bp completes its replication process within 38 minutes. Replication takes place
faster at the same time accurately. Any error will lead to mutation. However
replication errors are corrected by repair enzymes such as nucleases. Deoxy
nucleotide triphosphate acts as substrate and also provides energy for
polymerization reaction.
Replication begins at the initiation site called the site of ‘ origin of replication’ (ori). In prokaryotes, there is only one origin of replication, whereas in eukaryotes with giant DNA molecules, there can be several origins of replication (replicons). Since the two strands of DNA cannot be separated throughout at a time (due to large requirement of energy) the replication occurs within a small opening of the DNA helix called as replication fork. Unwinding of the DNA strand is carried out by DNA helicase. Thus, in one strand (template strand with polarity 3'→5') the replication is continuous and is known as the leading strand while in the other strand (coding strand with polarity 5'→3') replication is discontinuous, known as the lagging strand (Fig. 5.6). The discontinuously synthesized fragments of the lagging strand (called the Okazaki fragments) are joined by the enzyme DNA ligase.
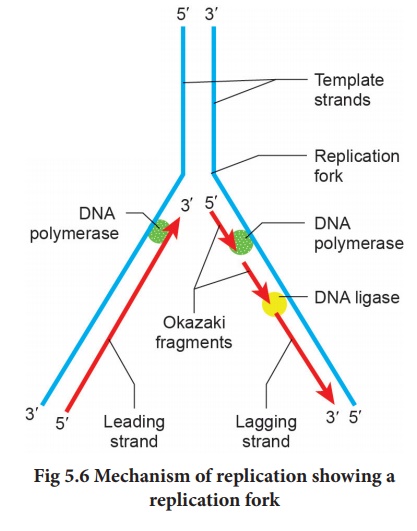
As they move away in
both directions, newly synthesized complementary nucleotides are paired with
the existing nucleotides on the parent strand and covalently bonded together by
DNA polymerase . Formation of new strand requires a primer (a short
stretch of RNA)for initiation. The primer produces a 3'-OH end on the sequence
of ribonucleotides, to which deoxy ribonucleotides are added. The RNA primer is
ultimately removed leaving a gap in the newly synthesized DNA strand. It is
removed from 5' end one by one by the exonuclease activity of DNA polymerase.
Finally, when all the nucleotides are in position, gaps are sealed by the
enzyme DNA ligase.
At the point of origin of repliction, the helicases and topoisomerases (DNA gyrase) unwind and pull apart the strands, forming a Y-Shaped structure called the replication fork. There are two replication forks at each origin. The two strands of a DNA helix have an antiparallel orientation. The enzyme DNA polymerase can only catalyse the addition of a nucleotide to the new strands in the 5'→3' direction, as it can only add nucleotides to the 3' carbon position.
Related Topics