Chapter: Essential Microbiology: Microbial Genetics
Genetic transfer in microorganisms
Genetic transfer in microorganisms
The various mechanisms of mutation described above result in an alteration in the genetic make-up of an organism. This can also occur by recombination, in which genetic material from two cells combines to produce a variant different to either parent cell.
In bacteria, this involves the transfer of DNA from one cell (donor) to another (recipient). Because transfer occurs between cells of the same generation (unlike the ge-netic variation brought about by sexual reproduction in eucaryotes, where it is passed to the next generation), it is sometimes referred to as horizontal transfer. There are three ways in which gene transfer can occur in bacteria, which we shall explore in the following sections.
Transformation
Transformation is the simplest of these, and also the first to have been described. We have already referred to the classic experiment of Fred Griffith in 1928, the first demonstration that genetic transfer can occur in bacteria. Griffith had previously demonstrated the existence of two strains of the bacterium Streptococcus pneumoniae, which is one of the causative agents of pneumonia in humans, and is also extremely virulent in mice. The S (smooth)-form produced a polysaccharide capsule, whilst the R (rough)-form did not. These formed recognisably different colonies when grown on a solid medium, but more importantly, differed in their ability to bring about disease in experimental animals. The R-form, lacking the protective capsule, was easily destroyed by the defence system of the host. Griffith observed the effects of injecting mice with bacterial cells of both forms; these are outlined below and in Figure 11.24. The results of experiments 1–3 were predictable, those of experiment 4 startlingly unexpected:
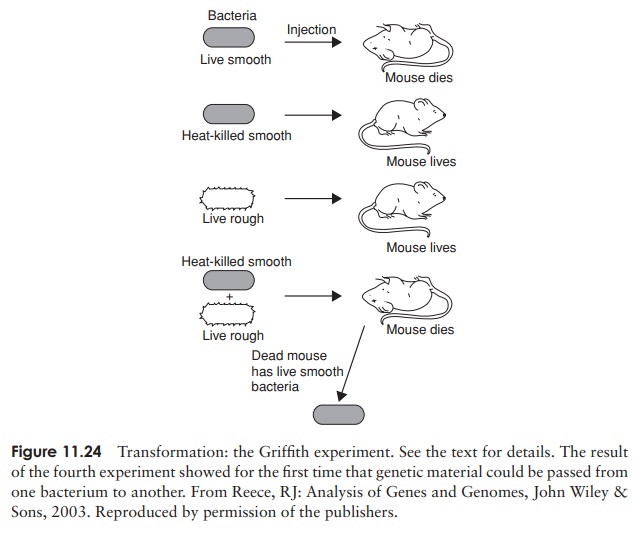
1 Mice injected with live cells of the R-form were unaffected.
2 Mice injected with live cells of the S-form died, and large numbers of S-form bacteria were recovered from their blood.
3 Mice injected with cells of the S-form that had been killed by heating at 60 ◦ C were unharmed and no bacteria were recovered from their blood.
4 Mice injected with a mixture of living R-form and heat-killed S-form cells died, and living S-form bacteria were isolated from their blood.
The S-form bacteria recovered from the mice in the crucial fourth experiment possessed a polysaccharide capsule like other S-forms, and, critically, were able to pass on thischaracteristic to subsequent generations. This finding went against the prevailing viewthat bacteria simply underwent binary fission, a completely asexual process involving no genetic transfer. Griffith deduced that some as yet unknown substance had passed from the heat-killed S-form cells to some of the living R-forms and conferred on them the ability to make capsules. Not long afterwards it was shown that this process of transformation could happen in the test tube, without the involvement of a host animal, and, as we have seen, it was eventually shown that Griffith’s ‘transforming principle’ was DNA.
How does transformation occur?
The uptake of foreign DNA from the environment is known to occur naturally in a num-ber of bacterial types, both Gram-positive and Gram-negative, by taking up fragments of naked DNA released from dead cells in the vicinity. Being linear and very fragile, the DNA is easily broken into fragments, each carrying on average around 10 genes. Transformation will only happen at a specific stage in the bacterial life cycle, when cells are in a physiological state known as competence. This occurs at different times in dif-ferent bacteria, but is commonly during late log phase. One of the reasons why only a low percentage of recipient cells become transformed is that only a small proportion of them are at any one time in a state of competence. The expression of proteins essential to the transformation process is dependant on the secretion of a competence factor.
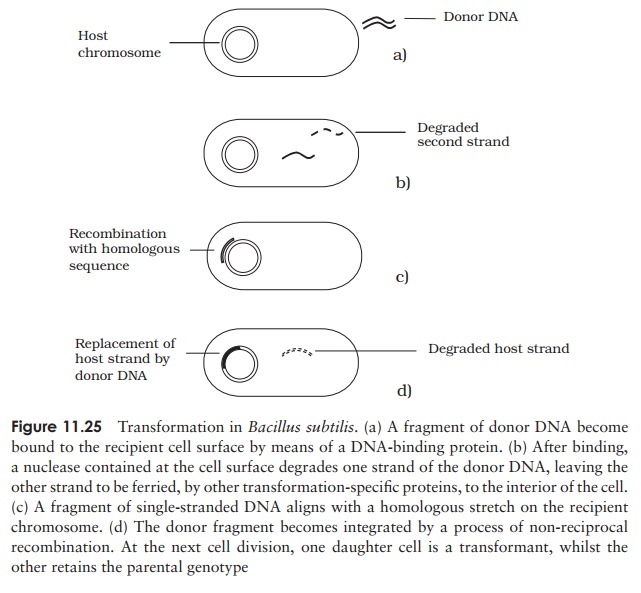
The exact mechanism of transformation varies somewhat according to species; the process for Bacillus subtilis is shown in Figure 11.25. Mere uptake of exogenous DNA is not enough to cause transformation; it must also be integrated into the host genome, displacing a single strand, which is subsequently degraded. Upon DNA replication and cell division, one daughter cell will inherit the parent genotype, and the othe the recombinant genotype. In the latter, there is a stable alteration of the cell’s genetic composition, and a new phenotype is expressed in subsequent generations.
Uptake of donor DNA from an unrelated species will not result in transformation; this is due to a failure to locate a homologous sequence and integrate into the host’s chromosome, rather than an inability to gain entry to the cell.
Induced competence
Transformation is not thought to occur naturally in E. coli, but, if subjected to certain treatments in the laboratory, cells of this species can be made to take up DNA, even from a completely unrelated source. This is done by effecting a state of induced or artificial competence, and by introducing the foreign DNA in a self-replicating vectormolecule, which does not depend on integration into the host chromosome. As we shall see, this is of enormous significance in the field of genetic engineering.
Conjugation
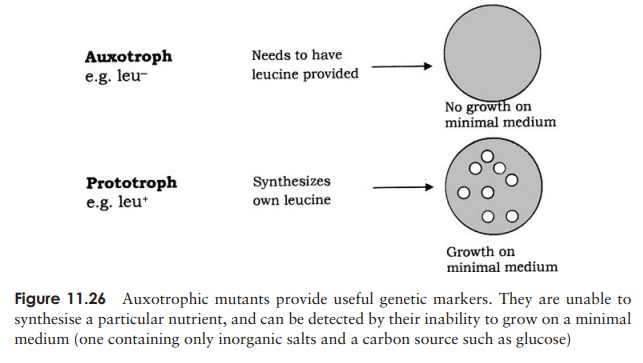
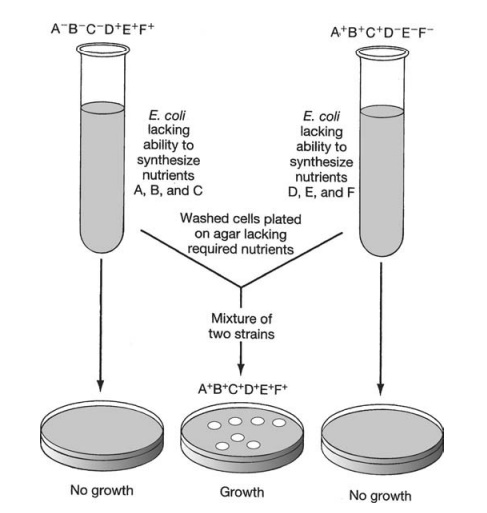
In 1946, Edward Tatum and Joshua Lederberg (the latter aged only 21 and still a student!) demonstrated a second form of genetic transfer between bacteria. These ex-periments involved the use of auxotrophic mutants, which have lost the ability to make a particular enzyme involved in the biosynthesis of an essential nutri-ent. These can be recognised experimentally by their inability to grow on a medium lacking the nutrient in question (Figure 11.26). The results of Tatum and Lederberg’s experiments suggested that a process was taking place in bacteria akin to sex in eucary-otes, in which there was a recombination of the cells’ genetic material (Figure 11.27). Transformation, as demonstrated by Griffith, could not explain their results, since the addition of a DNA-containing extract of one strain to whole cells of the other did not result in prototroph formation. This was confirmed in an ingenious experiment in which Bernard Davis showed that Tatum and Lederberg’s results were only obtained if direct cell-to cell contact was allowed (Figure 11.28).

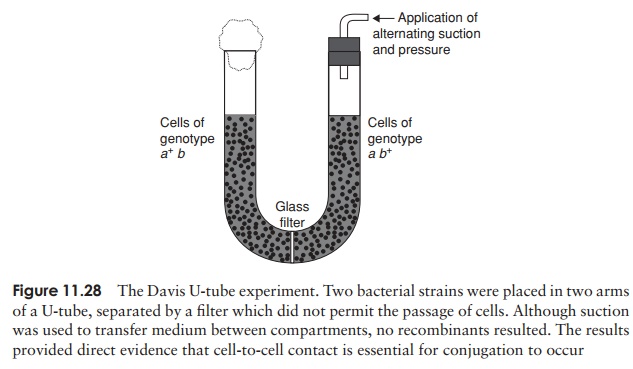
Gene transfer in conjugation is one way only
The process of conjugation was initially envisaged as a fusion of the two partner cells to give a diploid zygote, which subsequently underwent meiosis to give haploid offspring with modified genotypes. The work of William Hayes, however, showed that the devel-opment of colonies of recombinant cells was dependant on the survival of only one of the participating strains, the other strain being required only as a donor of DNA.
It became apparent that in E. coli, there are two distinct mating types, which became known as F+ and F− , depending on whether or not they possessed a plasmid called the F (fertility) plasmid. This contains some 30 or 40 genes responsible for its own replication and for the synthesis of a thread-like structure expressed on the cell surface called a sex pilus. In a mixture of F+ and F− cells, the sex pilus contacts an F− cell, then
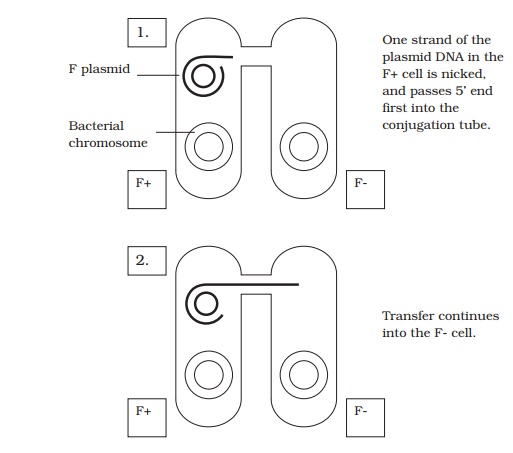
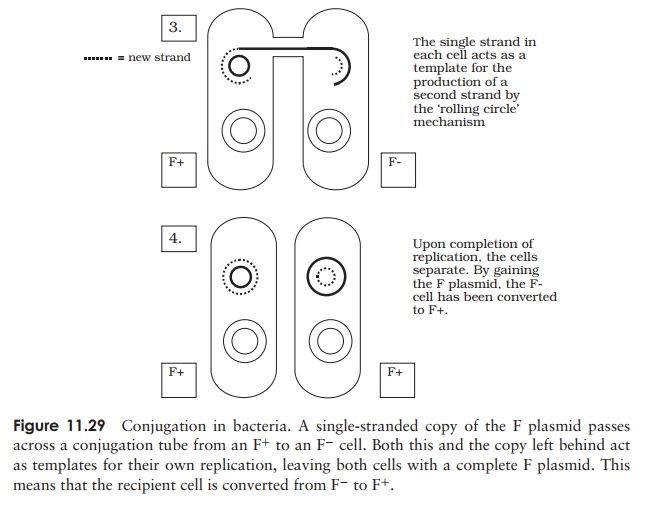
contracts, to pull the two cells together. A single strand of plasmid DNA is then passed across a channel made between the two cells, and enters the F− cell (Figure 11.29). This then serves as a template for the production of the complementary second strand, and similarly the single strand left behind in the donor cell is replicated, leaving us with a double-stranded copy of the F plasmid in both cells. By acquiring the F plasmid, the recipient F− cell has been converted to F+ .
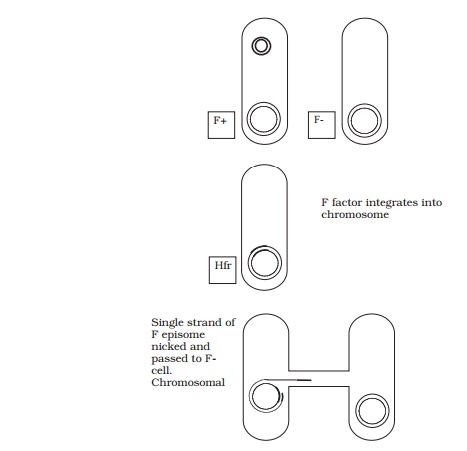
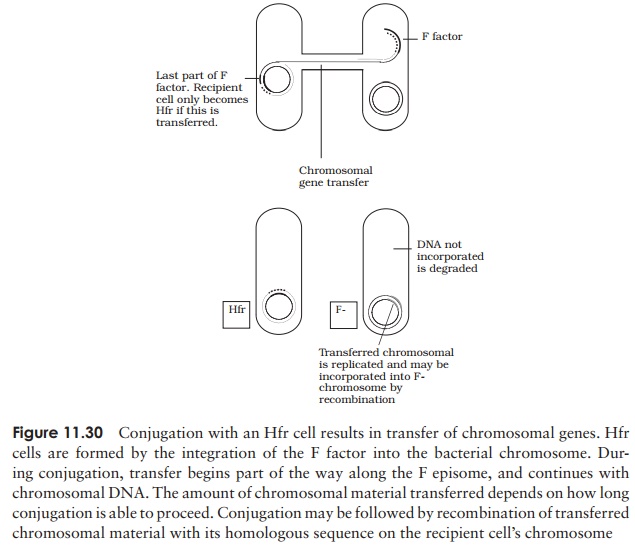
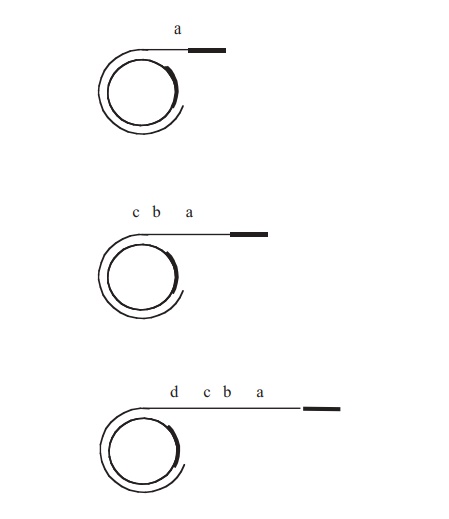
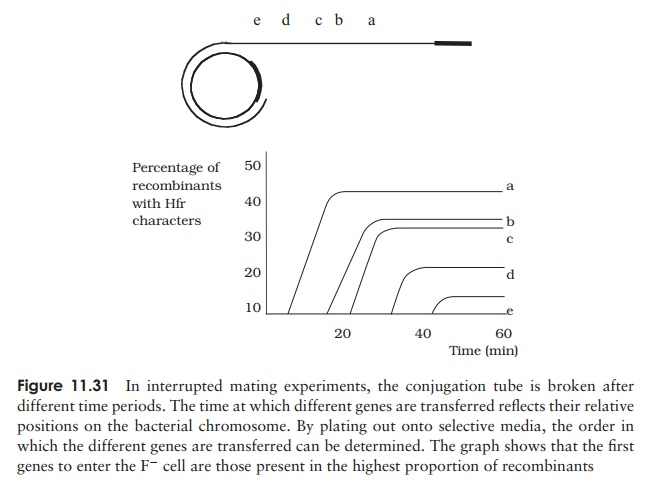
When conjugation involves a form of donor cell called Hfr (high frequency of recom-bination), genes from the main bacterial chromosome may be transferred. In these, the F plasmid has become integrated into the main bacterial chromosome; and thus loses its ability to replicate independently (Figure 11.30). It behaves just like any other part of the chromosome, although of course it still carries the genes for conjugation and pilus formation. When conjugation occurs, a single strand of Hfr DNA is broken within the F sequence, and is transferred in a linear fashion, carrying behind it chromosomal DNA. Recombination in the recipient cell results in the replacement of a homologous segment of DNA by the transferred material, which is then faithfully replicated in subsequent generations. If transfer is uninterrupted, a process which takes around 100 minutes in E. coli, eventually the remaining stretch of F plasmid will enter the F−cell, bringing upthe rear. The fragile nature of the pilus means, however, that transfer is rarely complete, and only a limited portion of the bacterial genome is transferred. This means that those F− cells receiving DNA from Hfr cells usually remain F− , unlike those in a cross with F+ cells, because the remainder of the F sequence is not transferred. It soon became clear that this phenomenon afforded a great opportunity to determine the relative positions of genes on the bacterial chromosome. This was done by interrupted mating experiments, in which the time allowed for conjugation is deliberately limited by mechanical break-age of the sex pili, and correlated with the phenotypic traits transferred to the recipient cells (Figure 11.31). By these means agenetic map of the bacterial chromosome could be developed.
The integration of the F plasmid into the bacterial chromosome is reversible; thus Hfr cells can revert to F+ . Excision of the integrated plasmid is not always precise, and sometimes a little chromosomal DNA is removed too. When this happens, the plasmid, and the cell containing it, are called F (‘F prime’); transfer of the plasmid to an F−cell takes with it the extra DNA from the host chromosome. The recipient genome thus becomes partially diploid (merodiploid), because it has its own copy, plus the ‘guest’ copy of certain genes.
Transduction
In the third form of genetic transfer in bacteria, bacteriophages act as carriers of DNA from one cell to another. In order to appreciate the way in which this is done, it is necessary to recall the sequence of events in phage replication cycles. (see Figure 10.11).
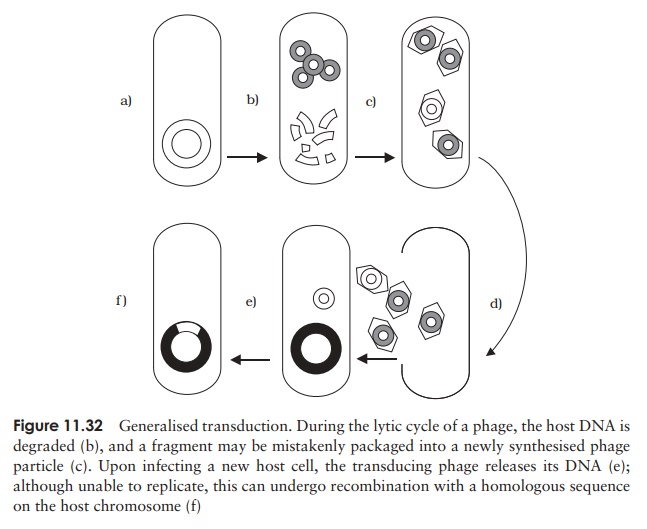
Generalised transduction occurs in virulent phages, that is, those with a lytic lifecycle. Sometimes, the enzymes responsible for packaging phage DNA into its protein coat package instead similarly sized fragments of degraded chromosomal DNA (Figure 11.32). Despite containing the wrong DNA, this transducing phage particle is still infec-tive, since this is dependant on its protein element. Thus following infection of another bacterial cell, the DNA can be incorporated by recombining with the homologous seg-ment in the recipient cell. Since any chromosomal fragment can be mistakenly packaged in this way (as long as it finds an area of homology), all genes are transferred at a similar (low) frequency.
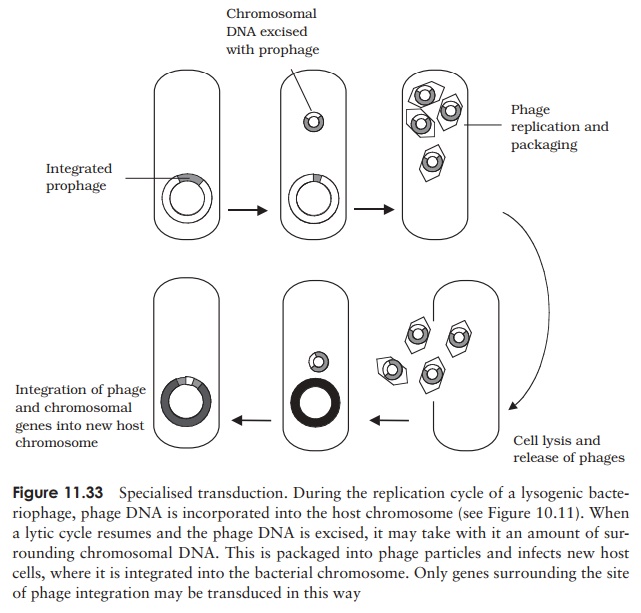
Specialised transduction results in a much higher efficiency of transfer for specificgenes, however it is limited to genes having a particular chromosomal location. Recall that in lysogenic life cycles, the phage DNA is integrated into the host chromosome, and later, perhaps after many rounds of cell division, excised again before re-entering a lytic cycle. If this excision does not happen precisely, some of the adjoining chromosomal DNA, carrying a gene or two, may be incorporated into the phage particle (we saw a similar mechanism in the case of F plasmid formation). Upon infecting another cell, the transduced genes would undergo recombination and become incorporated into the recipient’s chromosome (Figure 11.33). Although limited to genes in the vicinity of the lysogenic phage’s integration, this is a highly efficient form of transfer, since the genes become stably integrated into the host cell.
Transduction experiments, like those involving conjugation, can be used to determine the relative positions of genes on a bacterial chromosome.
Transposable elements
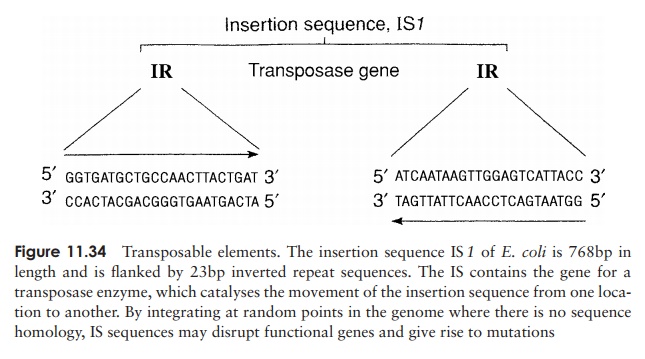
An unusual type of genetic transfer which takes place within an individual cell involves sequences of DNA called transposable elements. One type is known as aninsertionsequence (IS), a relatively short piece of chromosomal or plasmid DNA which containsa gene for the enzyme transposase (Figure 11.34). This recognises, cuts and re-ligates the insertion sequence anywhere in the bacterial genome. In so doing, it may interrupt a gene sequence, and thereby cause a mutation. Unlike re-combination events, no homology is required between the transposable element and the point at which it in-serts. This relocation of a transposable element from one place in the genome to another is termed conservativetransposition. In replicative transposition, the elementremains in its original position and a copy is made and inserted elsewhere in the genome. Insertion sequences are flanked by inverted sequences some 9–41bp in length, which are thought to be essential for the recognition of the sequence by the transposase.
Related Topics