Chapter: Essential Microbiology: Microbial Genetics
DNA replication
DNA replication
The elucidation of the structure of DNA by Watson and Crick in 1953 stands as one of the great scientific breakthroughs of the 20th century. An important implication of their model lay in the complementary nature of the two strands, as they commented themselves in their famous paper in Nature:
It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic material.
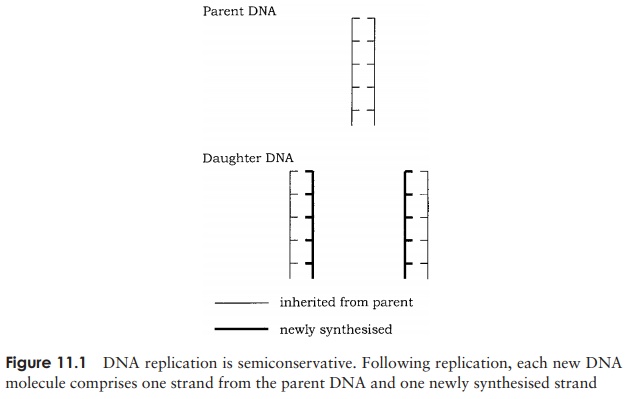
The mechanism they proposed became known as the semiconservative replication of DNA, so called because each daughter molecule comprises one parental strand and one newly synthesised strand (Figure 11.1). The parental double helix of DNA unwinds and each strand acts as a template for the production of a new complementary strand, with new nucleotides being added according to the rules of base-pairing. In 1957, 4 years after the publication of Watson and Crick’s model, Matthew Meselson and Franklin Stahl provided experimental proof of the semiconservative nature of DNA replication. The exact way in which replication takes place in procaryotic and eucaryotic cells differs in some respects, but we shall take as our model replication in E. coli, since it has been studied so extensively.
DNA replication in procaryotes
You may recall that bacteria multiply by a process of binary fission; before this occurs, each cell must duplicate its genetic information so that each daughter cell has a copy.
DNA replication involves the action of a number of specialised enzymes:
· Helicases
· DNA topoisomerases
· DNA polymerase I
· DNA polymerase III
· DNA primase
· DNA ligase.
Replication begins at a specific sequence called an ori-gin of replication. The two strands of DNA are caused toseparate by helicases (Figure 11.2), while single-strandedDNA binding proteins (SSB) prevents them rejoining.Opening out part of the double helix causes increased tension (supercoiling) elsewhere in the molecule, which is relieved by the enzyme DNA topoiosomerase (some-times known as DNA gyrase). As the ‘zipper’ moves along, and more single stranded DNA is exposed,DNApolymerase III adds new nucleotides to form a comple-mentary second strand, according to the rules of base-pairing. DNA polymerases are not capable of initiating the synthesis of an entirely new strand, but can only extend an existing one. This is because they require a free 3 -OH group onto which to attach new nucleotides. Thus, DNA polymerase III can only work in the 5 –3direction. A form of RNA polymerase called DNA pri-mase synthesises a short single strand of RNA, whichcan be used as aprimer by the DNA polymerase III. (Figure 11.2).
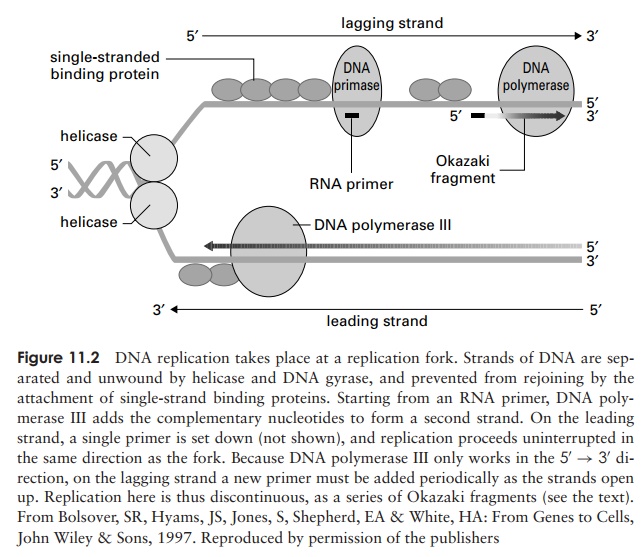
When replication occurs, complementary nucleotides are added to one of the strands (the leading strand) in a continuous fashion (Figure 11.2). The other strand (thelagging strand), however, runs in the opposite polarity; so how is a complementary sequence synthesised here? The answer is that DNA polymerase III allows a little unwinding to take place and then, starting at the fork, works back over from a new primer, in the 5 –3 direction. Thus, the second strand is synthesised discontinuously, in short bursts, about 1000–2000 nucleotides at a time. These short stretches of DNA are called Okazakifragments, after their discoverers.On the lagging strand, a new RNA primer is needed at the start of every Okazaki fragment. These short sequences of RNA are later removed by DNA polymerase I, which
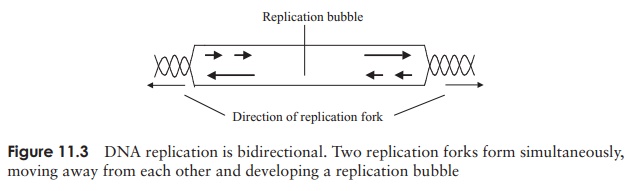
then replaces them with DNA nucleotides. Finally, the fragments are joined together by the action of DNAligase. Replication is bidirectional (Figure 11.3), withtwo forks moving in opposite directions; when they meet, the whole chromosome is copied and replication is complete* .
What happens when replication goes wrong?
It is clearly important that synthesis of a complementary second strand of DNA should occur with complete accuracy, but occasionally, a non-complementary nucleotide is inserted; this may happen as frequently as once in every 10 000 nucleotides. Cells, however, are able to have a second attempt to incorporate the correct base because of the proofreading activity of the enzymes DNA polymerase I and III. These are able to cut out the ‘wrong’ nucleotide and replace it with the correct one. As a result of this monitoring, mistakes are very rare; they are thought to occur at a frequency of around one in every billion (109) nucleotides copied. Mistakes that do slip through the net result in mutations.
DNA replication in eucaryotes
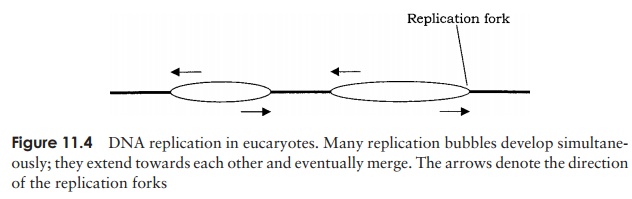
In eucaryotic organisms, the DNA is linear, not circular, so the process of replication differs in some respects. Genome sizes are generally much bigger in eucaryotes, and replication much slower, so numerous replication forks are active simultaneously on each chromosome. Replication takes place bidirectionally, creating numerous replica-tion bubbles (Figure 11.4) which merge with one another until the whole chromosome has been covered.
Related Topics