Chapter: Biotechnology Applying the Genetic Revolution: Protein Engineering
Engineered Binding Proteins
ENGINEERED
BINDING PROTEINS
Making drugs specific for a particular organ can eliminate many unwanted side effects. One way of achieving this is to attach the drug to a reagent that recognizes proteins specific to particular tissues. As noted in Previews Pages, antibodies are the most widely used reagents for binding specific target proteins. However, antibodies require disulfide crosslinks to function, and these are often hard to maintain during large-scale manufacture. Some researchers have therefore been seeking alternatives to antibodies.
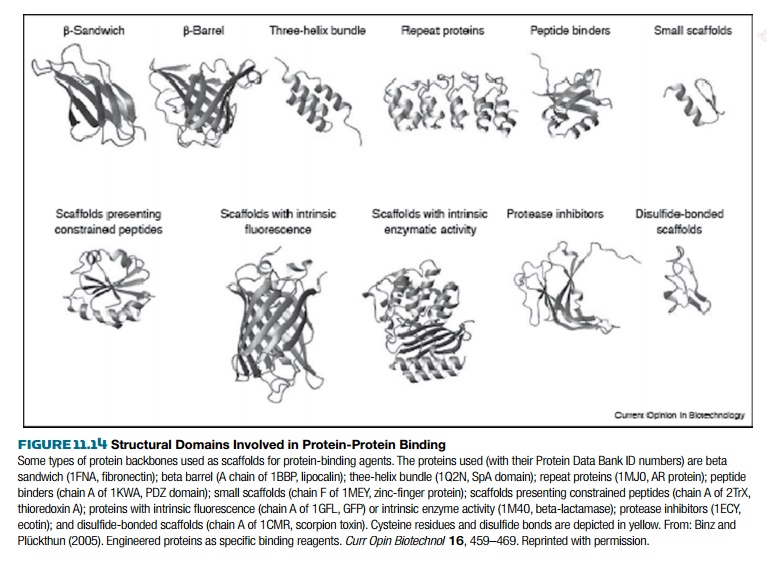
Most research into finding
nonantibody binding partners has focused on certain protein structural domains.
Because many different proteins have been crystallized and their structures
have been determined, many different binding domains can be compared. The
binding domains of one family of proteins share the same general structure,
such as a β-barrel or three-helix bundle
(Fig. 11.14). To generate novel binding domains, a binding protein with a known structure is chosen and
the amino acid residues associated with binding are identified. The binding
protein is modified by mutation of these residues and then screened for new
binding partners. It is hoped that the targeted directed evolution approach
will find new, more easily isolated proteins for targeting drugs to specific
target cells within our bodies.
Related Topics