Chapter: Biotechnology Applying the Genetic Revolution: Transgenic Animals
Applications of RNA Interference in Transgenics
APPLICATIONS
OF RNA INTERFERENCE IN TRANSGENICS
RNA interference covers
several related phenomena. The mechanisms are still only partly understood, but
involve the formation of double-stranded RNA (dsRNA). This is not normally made
by either prokaryotic or eukaryotic cells. Consequently it is regarded by
eukaryotic cells as evidence of viral activity and therefore is degraded.
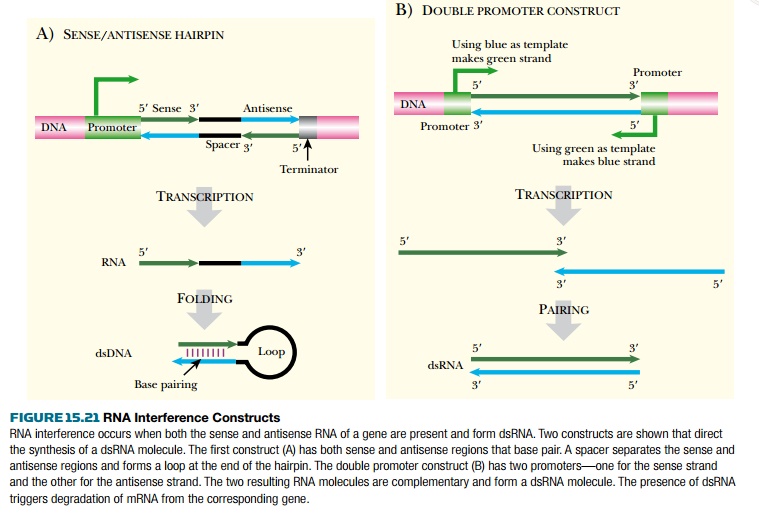
When both sense and antisense RNA are present in eukaryotic cells, the formation of dsRNA is followed by a massive drop in expression of the corresponding gene. This effect is due to degradation of the mRNA. Therefore the dsRNA must correspond to the exon sequences of the target gene. Only a few molecules of dsRNA are needed to trigger this effect. RNA interference is the favored procedure for gene inactivation in Caenorhabditis elegans and is also effective in other animals and plants.
RNA interference can be
triggered by adding RNA directly or by expression of the corresponding transgenic
DNA constructs. A construct with neighboring sense and antisense sequences will
produce hairpin RNA (Fig. 15.21). Alternatively it is possible to use a single
transgene flanked by two promoters, one pointing in each direction.
Homologous cosuppression is a type of RNAi characteristic of plants,
though occasionally seen in other
organisms. Here the presence of multiple copies of a transgene decreases the
expression of related host genes (and of the transgenes themselves, too).
Occasionally, single copies of transgenes that are highly homologous to an
endogenous host gene may be sufficient to trigger cosuppression. The mechanism
varies somewhat from case to case, but results in formation of dsRNA, which
then triggers RNA interference. Transgenes carried by RNA viruses are
especially effective at causing homologous cosuppression, although only if the
virus is still capable of replication.
Related Topics