Chapter: Biotechnology Applying the Genetic Revolution: Immune Technology
Antibody Engineering
ANTIBODY
ENGINEERING
Natural antibodies consist of
an antigen binding site joined to an effector region that is responsible for
activating complement and or binding to immune cells. From a biotechnological
viewpoint, the incredibly high specificity with which antibodies bind to a
target protein is useful for a variety of purposes. Consequently, antibody
engineering uses the antigen binding region of the antibody. These are
manipulated and are attached to other molecular fragments.
To separate an antigen
binding site from the rest of the antibody, gene segments encoding portions of
antibody chains are subcloned and expressed in bacterial cells. Bacterial
signal sequences are added to the N terminus of the partial antibody chains,
which results in export of the chains into the periplasmic space. Here the VH
and VL domains fold up correctly and form their disulfide bonds. The antibody
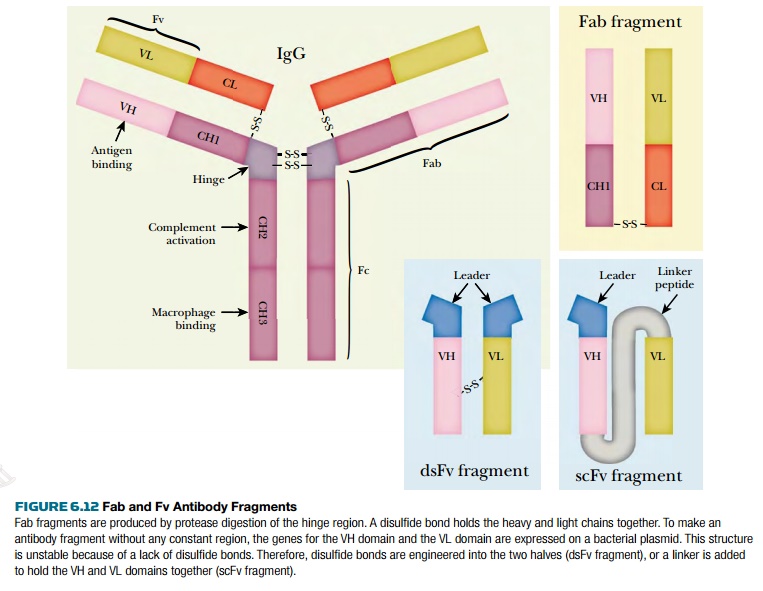
fragments used include Fab, Fv, and single-chain
Fv (scFv) (Fig. 6.12). In a Fab
fragment, an interchain disulfide bond holds the two chains together. However, the Fv fragment lacks this region of the
antibody chains and thus is less stable. This led to development of the
single-chain Fv fragment in which the VH and VL domains are linked together by
a short peptide chain, usually 15 to 20 amino acids long. This is introduced at
the genetic level so that a single artificial gene expresses the whole
structure (VH-linker-VL or VL-linker-VH). A tag sequence (such as a His6-tag or
FLAG-tag) is often added to the end to allow detection and purification. Such
an scFv fragment is quite small, about 25,000 in molecular weight.
Such scFv fragments are attached
to various other molecules by genetic engineering. The role of the scFv
fragment is to recognize some target molecule, perhaps a protein expressed only
on the surface of a virus-infected cell or a cancer cell. A variety of toxins,
cytokines, or enzymes may be attached to the other end of the scFv fragment, to
provide the active portion of the final recombinant antibody. In principle,
this approach provides a way of delivering a therapeutic agent in an extremely
specific manner. At present the clinical applications of engineered antibodies
are under experimental investigation.
The antigen binding regions
used in antibody engineering may be derived from characterized monoclonal
antibodies. Alternatively, a library of DNA segments encoding V-regions may be
obtained from a pool of B cells obtained from an animal or human blood sample.
Such a library should in theory contain V-regions capable of recognizing any
target molecule. Using a human source avoids the necessity for the complex
humanization procedures described earlier. However, in this case it is
necessary to screen the V-region library for an antibody fragment that binds to
the desired target molecule. This may be done by the phage display procedure
outlined. The library of V-region constructs is expressed on the surface of the
phage, and the target molecule is attached to some solid support and used to
screen out those phages carrying the required antibody V region.
Related Topics