Chapter: Plant Biochemistry: A plant cell has three different genomes
Viruses are present in most plant cells
Viruses are present in most plant cells
With the exception of meristematic cells, almost all other plant cells are infected by viruses. In most cases, viruses do not kill their host since they depend on the host’s metabolism for reproduction. The viruses encode only a few special proteins and use the energy metabolism and the biosynthetic capacity of the host cell to multiply. This often weakens the host plant and lowers the yield of virus-infected cultivars. Infection by some viruses can lead to the destruction of the entire crop. Courgettes and melons are extremely susceptible to the cucumber mosaic virus. In some provinces of Brazil, 75% of the orange trees were destroyed within 12 years by the Tristeza virus.
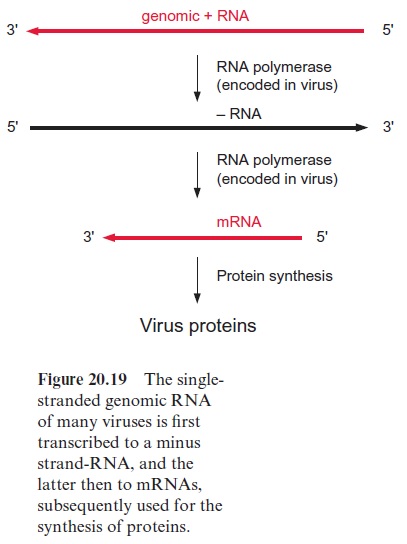
The virus genome consists of RNA or DNA surrounded by a protein coat. The majority of plant virus genomes consist of a single-stranded RNA, called the plus RNA strand. In some viruses (e.g., brome mosaic virus, which infects certain cereals), the plus RNA strand exhibits the characteristics of an mRNA and is therefore translated by the host cell. In other viruses (e.g., the tobacco mosaic virus (TMV)), the plus RNA strand is first transcribed to a complementary minus RNA strand and the latter then serves as a template for the formation of mRNAs (Fig. 20.19). The translation products of these mRNAs encompass replicases, which catalyze the replication of the plus and minus RNAs, movement proteinsthat enable the spreading of the viruses from cell to cell , and coat proteins for packing the virus DNA. Normally, a virus infiltrates a cell through wounds, which, for instance, have been caused by insects, such as aphids feeding on the plant . Once viruses have entered the cell, their movement proteins widen the plasmodesmata between single cells to allow virus passage and spreading over the entire symplast .
The retrovirus genome consists also of a single-stranded RNA, but in this case, when the cell has been infected, the RNA is transcribed into DNA by a reverse transcriptase. The DNA is integrated in part into the nuclear genome. So far, infections by retroviruses are known only in animals.
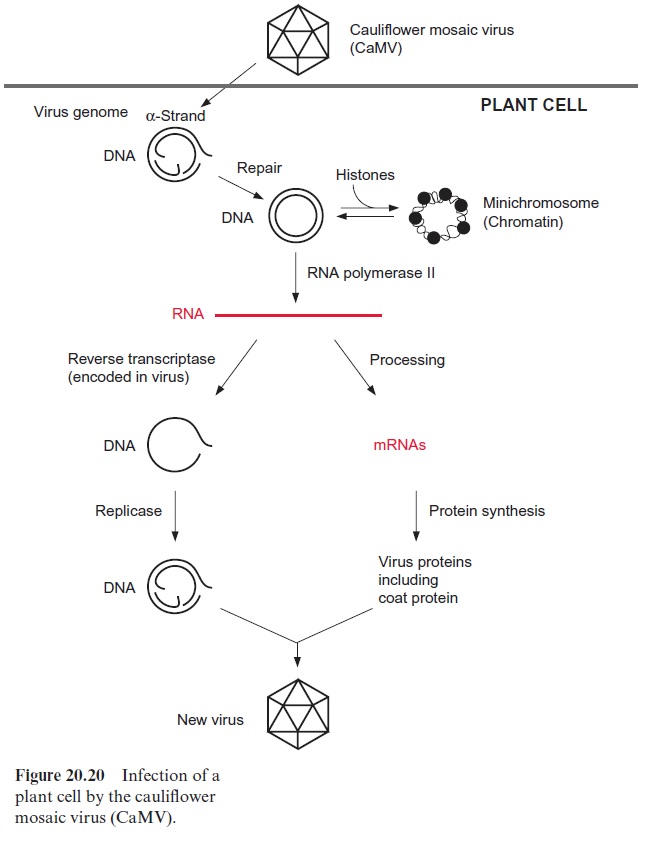
The cauliflower mosaic virus (CaMV), which causes pathogenic changes in leaves of cauliflower and related plants, is somewhat similar to a retrovi rus. The genome of the CaMV consists of a double-stranded DNA of about 8 kbp, with gaps in it (Fig. 20.20). When a plant cell is infected, the virus loses its protein coat and the gapped DNA strands are repaired by enzymes of the host. The virus genome acquires a double helical structure and forms in the nucleus a chromatin-like aggregate with the histones. This permits the viral genome to stay in the nucleus as a mini-chromosome. The viral genome possesses promoter sequences that are similar to those of nuclear genes, such as TATA box and a CAAT box as well as enhancer elements. The virus promoter (35S promoter) is recognized by the RNA polymer ase II of the host cell and is transcribed at a high rate. The precursor tran script is subsequently processed into individual mRNAs, which encode the synthesis of six virus proteins, including the coat protein and the reverse transcriptase. The strong CaMV 35S promoter is often used as a promoter for the expression of foreign genes in transgenic plants .
The transcript synthesized by RNA polymerase II is also reverse tran scribed by the virus-encoded reverse transcriptase into DNA and, after syn thesis of the complementary strand, packed as double-stranded DNA into a protein coat. This mature virus is now ready to infect other cells.
Retrotransposons are degenerated retroviruses
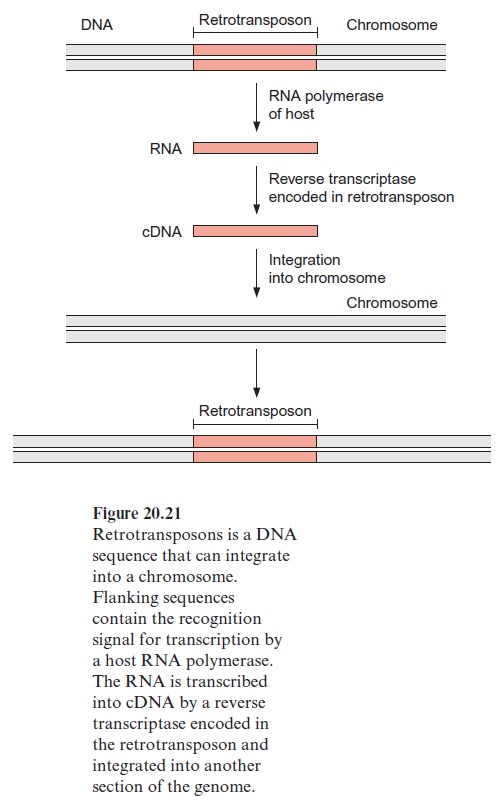
Besides the transposons there is another class of mobile elements, which are derived from the retroviruses. They do not jump out of a gene like the trans posons but just multiply. At both ends of the retrotransposons sequences are present, which carry signals for the transcription of the retrotransposon DNA by the host RNA polymerase. The retrotransposon RNA does not encode the coat protein, but the reverse transcriptase, which is homologous to the retrovirus enzyme and transcribes the retrotransposon RNA into DNA (Fig. 20.21). This DNA is then inserted into another site of the plant genome. It is assumed that these retrotransposons originate from retrovi ruses that have lost the ability to synthesize the protein coat. Several differ ent retrotransposons containing all the constituents for their multiplication have been found in Arabidopsis, but so far an insertion of a retrotransposon into a gene ofArabidopsis has never been monitored. About 0.1% of the genome of Arabidopsis consists of these retrotransposons, suggesting that these actually multiply, albeit at a slow rate.
Related Topics