Chapter: Plant Biochemistry: A plant cell has three different genomes
Plastids possess a circular genome
Plastids possess a circular genome
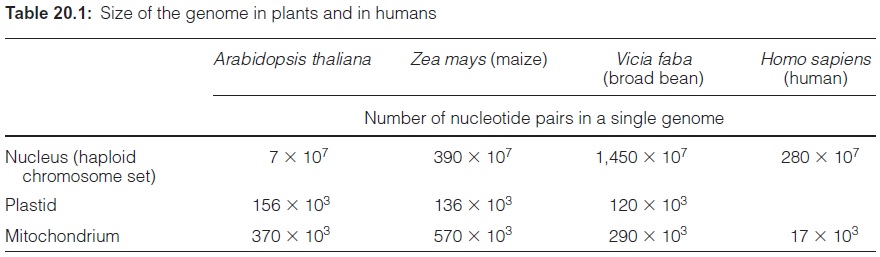
Many arguments support the hypothesis that plastids have evolved from prokaryotic endosymbionts. The circular genome of the plas tids is similar to the genome of the prokaryotic cyanobacteria, although much smaller. The DNA of the plastid genome is named ctDNA (chloroplast) or ptDNA (plastid). In the majority of the plants investigated so far, the circular plastid genome has the size of 120 to 160 kbp. Depending on the plant, this is only 0.001% to 0.1% of the size of the nuclear genome (Table 20.1), but the cell contains many copies of the plastid DNA, because each plastid contains many genome copies. In young leaves, the number of ctDNA molecules per chloroplast is about 100, whereas in older leaves, it is between 15 and 20. Furthermore, a cell contains a large number of plastids, a mesophyll cell, for instance 20 to 50. Thus, despite the small size of the plastid genome, the plastid DNA can amount to 5% to 10% of the total cellular DNA.
The first complete analysis of the nucleotide sequence of a plastid genome was carried out in 1986 by the group of Katzuo Shinozaki in Nagoya with chloroplasts from tobacco and by Kanji Ohyama in Kyoto with chloroplasts from the liverwort Marchantia polymorpha. Although the two investigated plants are very distantly related, their plastid genomes are rather similar in gene composition and arrangements. Obviously, the plastid genome has changed little during recent evolution. Present analysis of the DNA sequence of plastid genes from many plants supports this notion.
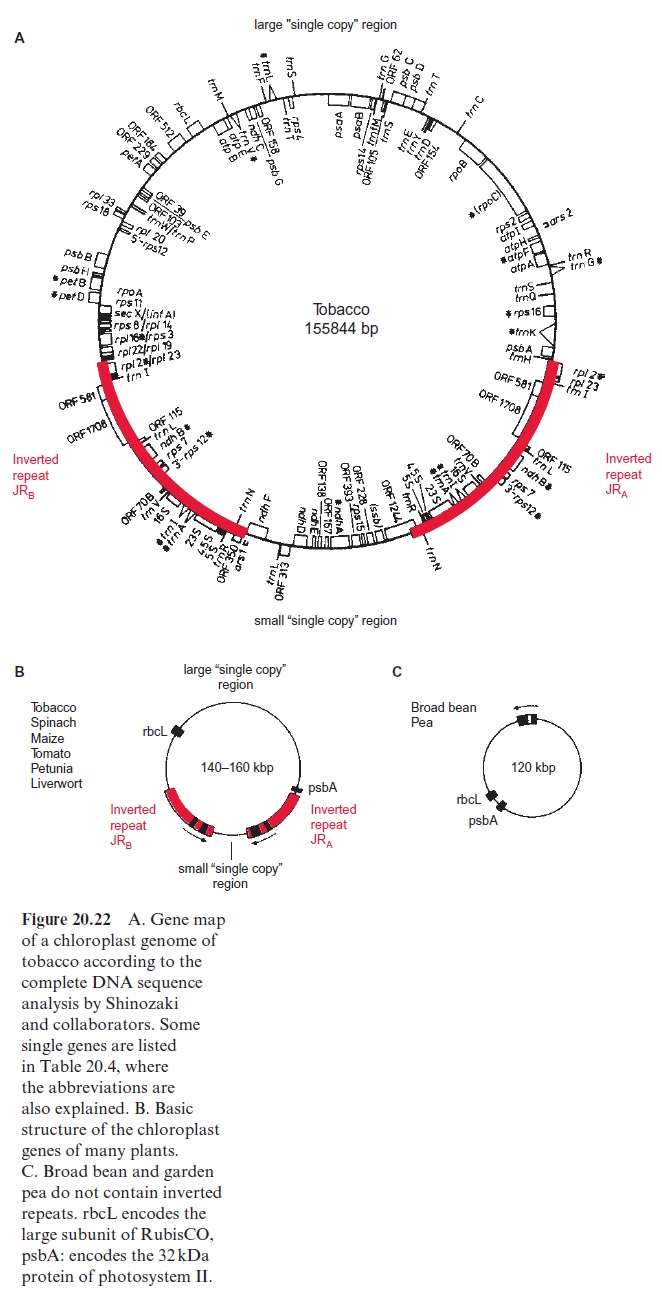
Figure 20.22A shows a complete gene map of the chloroplast genome of tobacco and Figure 20.22B shows schematic representations of the plastid genomes of other plants. The plastid genome of most plants contains so-called inverted repeats (IR), which divide the remaining genome into a large or small single copy region. The repeat IRA and IRB each encode the genes for the four ribosomal RNAs as well as the genes for some transfer RNAs, and the repeat sizes vary from 20 to 50 kbp. These inverted repeats are not found in the plastid genomes of pea, broad bean, and other legumes (Fig. 20.22C), where the inverted repeats probably have been lost during the course of evolution. On the remainder of the genome (single-copy region), genes are present usually only in a single copy.
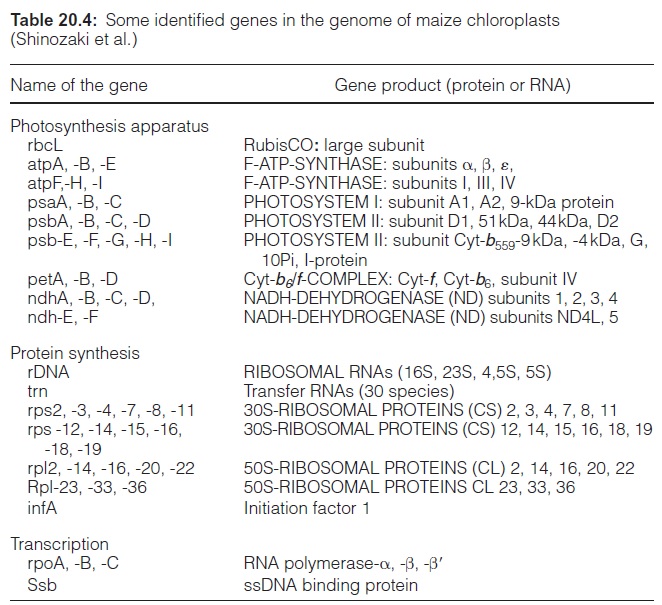
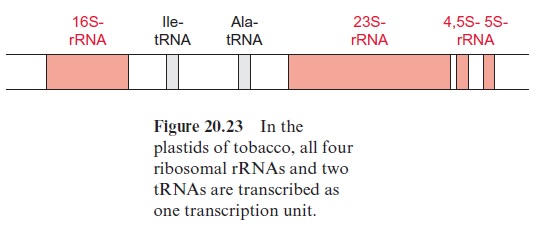
Analysis of the ctDNA sequence of tobacco revealed that the genome encodes 122 genes (146 if the genes of each of the two inverted repeats are counted) (Table 20.4). The gene for the large subunit of ribulose bisphos phate carboxylase/oxygenase (RubisCO) is located in the large single-copy region, whereas the gene for the small subunit is present in the nuclear genome. The single-copy region of the plastid genome also encodes six subunits of F-ATP synthase, whereas the remaining genes of F-ATP synthase are encoded in the nucleus. Also encoded in the plastid genome are subunits of photosystem I and II, of the cytochrome-b6/f complex, and of an NADH dehydrogenase (which also occurs in mitochondria,), and furthermore, proteins of plastid protein synthe sis and gene transcription. Some of these plastid structural genes contain introns. In addition, there are putative genes on the genome with so-called open reading frames (ORF), which, like the other genes, are bordered by a start and a stop codon, but where the encoded proteins are not yet known. The plastid genome encodes only a fraction of plastid proteins, as the majority is encoded in the nucleus. It is assumed that many genes of the original endosymbiont have been transferred during evolution to the nucleus, but there also are indications for gene transfer between the plas tids and the mitochondria .
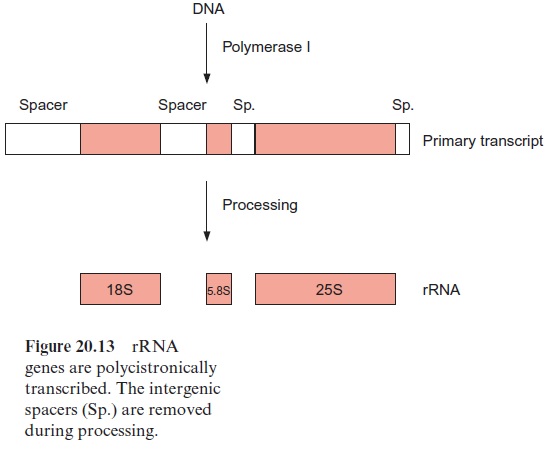
All four rRNAs, which are constituents of the plastid ribosome (4.5S-, 5S-, 16S-, and 23S-rRNA) are encoded in the plastid genome. The plastid ribosomes (sedimentation constants 70S) are smaller than the eukaryotic ribosomes (80S) present in the cytosol, but are similar in size to the ribosomes of bacteria. As in bacteria, these four rRNAs are encoded in the plastid genome in one transcription unit (polycistronic transcription) (Fig. 20.23). Between the 16S and 23S DNA a large spacer is situated (intergenic spacer), which encodes the sequence for one or two tRNAs. In total, about 30 tRNAs are encoded in the plastid genome. Additional tRNAs needed for transcrip tion in the plastids are encoded in the nucleus.

The transcription apparatus of the plastids resembles that of bacteria
In the plastids two types of RNA polymerases are active, of which only one is encoded in the plastid genome and the other in the nucleus:
1. The RNA polymerase encoded in the plastids enables the transcrip tion of plastid genes for subunits of the photosynthesis complex. This RNA polymerase is a multienzyme complex resembling that of bacteria. In contrast to the RNA polymerase of bacteria, the plastid enzyme is insensitive to rifampicin, a synthetic derivative of an antibiotic fromStreptomyces.
2. The plastid RNA polymerase, which is encoded in the nucleus, is derived from the duplication of mitochondrial RNA polymerase. This “imported” RNA polymerase is homologous to RNA polymerases from bacteriophages. The nucleus-encoded RNA polymerase transcribes the so-called housekeeping genes in the plastids. These are the genes that have general functions in metabolism, such as the synthesis of rRNA or tRNA.
As in bacteria, many plastid genes contain a box 10 bp upstream from the transcription start with the consensus sequence TATAAT and at 35 bp upstream a further promoter site with the consensus sequence TTGACA. Some structural genes are polycistronically transcribed, which means that several are in one transcription unit and are transcribed together as a large primary transcript. Polycistronic transcription often occurs with bacterial genes. In some cases, the primary transcript is subsequently processed by ribonucleases of which many details are still not known (see Figure 20.13)
Related Topics