Chapter: Plant Biochemistry: A plant cell has three different genomes
The mitochondrial genome of plants varies largely in its size
The mitochondrial genome of plants varies largely in its size
In contrast to animals, plants possess a very large mitochondrial (mt) genome. In Arabidopsis it is 20 times, and in melon 140 times larger than in humans (Table 20.5). The plant mitochondrial genome also encodes more genetic information: the number of encoding genes in a plant mt-genome is about seven times higher than in humans.
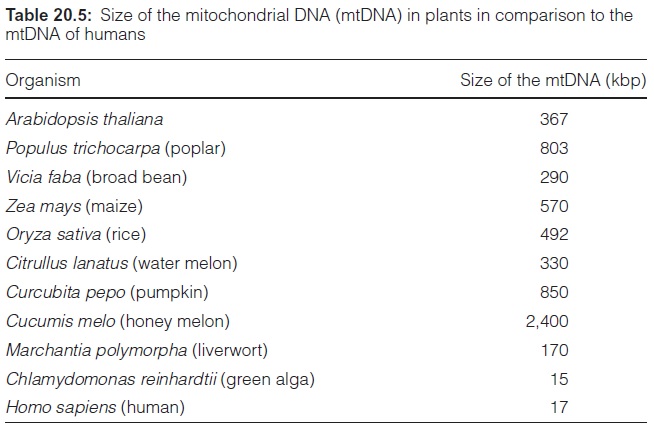

The size of the mt-genome varies largely in higher plants, even within a family. Citrullus lanatus (330 kbp), Curcubita pepo (850 kbp), and Cucumis melo (2,400 kbp), listed in Table 20.5, all belong to the family of the Curcubitaceae (squash plants). In most plants, however, the size of the plas tid genome is relatively constant at 120 to 160 kbp.
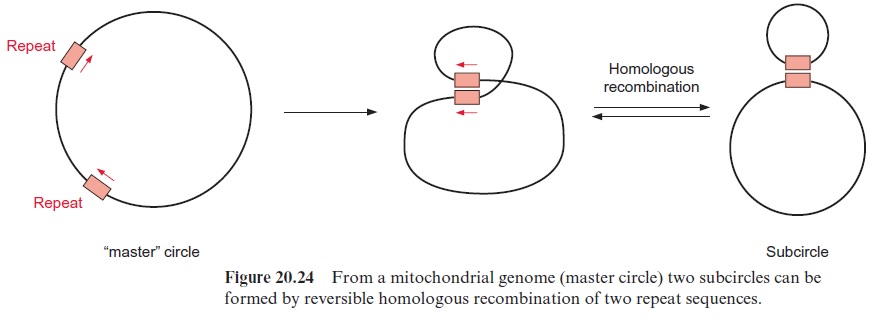
The mitochondrial genome in plants often consists of one large circular DNA molecule and several smaller ones. In some mitochondrial genomes, this partitioning may be permanent, but in many cases the fragmentation of the mt-genome seems to be derived from homologous recombination of repetitive elements (e.g., maize contains six such repeats). Figure 20.24shows how an interaction of two repeats can lead by homologous recom-bination to a fragmentation of a DNA molecule. The 570 kbp mt-genome of maize is present as a master circle as well as up to four subcircles (Fig. 20.25). The homologous recombination of DNA molecules can also form larger units. This may explain the large variability in the size of the mito chondrial genomes in plants.
The number of mitochondria in a plant cell can range between 50 and 2,000, with each mitochondrion containing 1 to 100 genomes.
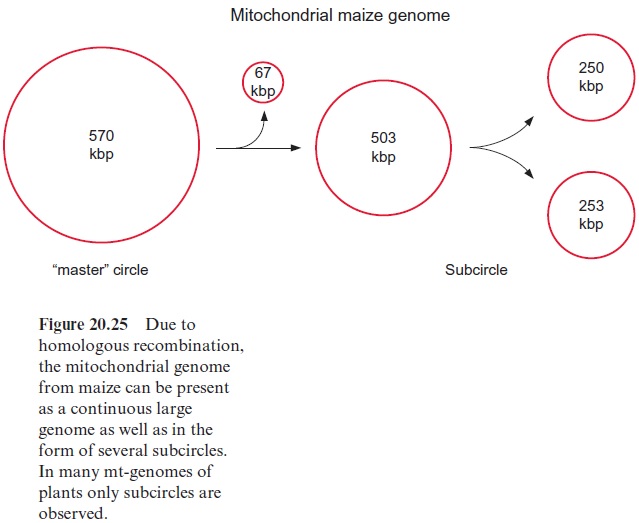
In animals and in yeast, the mtDNA is normally circular, like the bacte rial DNA. Also plant mtDNA is thought to be circular. This is undisputed for the small mtDNA molecules (subcircles, Fig. 20.25), but it remains unclear whether this circular structure also generally applies to the master mtDNA. There are indications that the master genome can also occur as open strands.
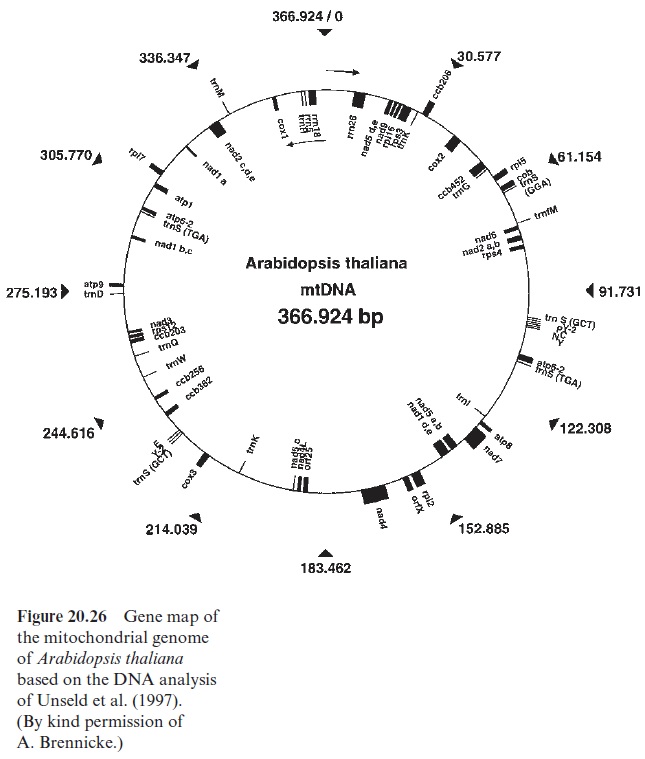
Figure 20.26 shows the complete gene map of the mt-genome of Arabidopsis thaliana, and Table 20.6 summarizes the genes in the mt-genome of a higher plant. A comparison with the number of genes in the plastid genome (Table 20.4) shows that the mt-genome of the plant, although usually much larger than the plastid genome, encodes much fewer genes. The relatively small informa tion content of the mt-genome in relation to its size is due to a high content of repetitive sequences, which probably are derived from gene duplication. The mt-genome contains much DNA with no recognizable function, termed junk DNA, which has accumulated in the mt-genome during evolution. Some of this junk DNA has its origin in plastid DNA and some in nuclear DNA. The mitochondrial genome, like the nuclear genome, apparently tolerates a large portion of apparently senseless sequences and passes these on to following gen erations. It is interesting that a large part of mtDNA is transcribed. About 30% of the mt-genome of Brassica rapa (218 kbp) is transcribed, as is that of the six times larger genome of Cucumis melo (2,400 kbp). Why so many transcripts are synthesized is obscure, considering that in the two mt-genomes mentioned above, the total number of the encoded proteins, tRNAs, and rRNAs amounts only to about 60.
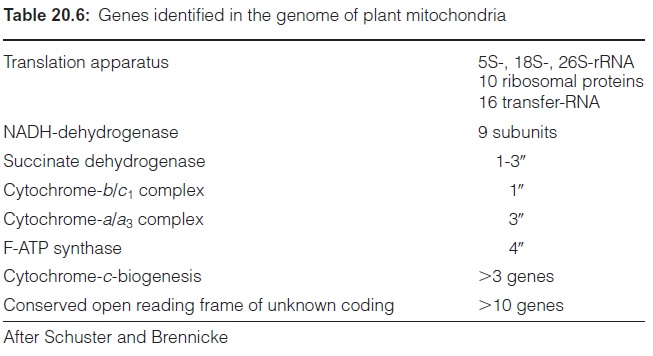

The mitochondrial genome encodes elements of the translation machin ery, including three rRNAs, about 16 tRNAs, and about 10 ribosomal proteins. These components of the translation apparatus are involved in the synthesis of various hydrophobic membrane proteins, which are also encoded in the mt-genome, e.g., some subunits of the respiratory chain , of F-ATP synthase , and at least three enzymes of cytochrome-c synthesis. About 95% of the mitochondrial proteins, including most subunits for the respiratory chain and F-ATP syn thase, as well as several tRNAs, are encoded in the nucleus. Considering that mitochondria have derived from endosymbionts, it must be assumed that the largest part of the genetic information of the endosymbiont genome has been transferred to the nucleus. Such gene transfers occur quite often in plants; the gene content in the mitochondria can vary between the different species. Moreover, genes apparently can pass from the plastids to the mitochondria. Delineated from their nucleotide sequences, several mt tRNA genes seem to originate in the plastid genome.
The promoters of plant mitochondrial genes are heterogeneous. The sequences signaling the start and end of transcription are quite variable, even for the genes within the same mitochondrion. Most likely several mtRNA polymerases are present in plant mitochondria, similarly as in plastids. Mitochondrial transcription factors have not yet been unequivocally charac terized. Most of the mitochondrial genes are transcribed monocistronically.
Mitochondrial RNA is corrected after transcription via editing
A comparison of the amino acid sequences of proteins encoded in mito chondria with the corresponding nucleotide sequences of the encoding genes revealed strange discrepancies: the amino acid sequences did not cor respond to the DNA sequences of the genome according to the universal genetic code. At sites of the DNA sequence where, according to the protein sequence a T was to be expected, a C was found, and sometimes vice versa. More detailed studies showed that the transcription of mtDNA yielded an mRNA sequence that would not translate into the “correct” (expected) protein. It was all the more astonishing to discover that this “incorrect” mRNA subsequently is processed in the mitochondria by several replace ments of C by U, but sometimes also of U by C, until the correct mRNA is reconstructed as a template for synthesizing the proper protein. This proc ess is called RNA editing.
Subsequent editing of the initially incorrect mRNA to the correct, translatable mRNA is not an exception taking place only in some exotic genes, but is the rule for the mitochondrial genes of higher plants. In some mRNAs produced in the mitochondria, 40% of the C is replaced by U in the editing process. Mitochondrial tRNAs are also edited in this way. The question arises whether, by differences in the editing, a structural gene can be translated into different proteins. Now and again mitochondrial proteins were found which had been translated from only partially edited mRNA. Since these proteins normally are nonfunctional, they are probably degraded rapidly.
RNA editing was first shown in the mitochondria of trypanosomes, the unicellular pathogen of sleeping sickness. It also occurs in mitochondria of animals and, in a few cases, has also been found in plastids.
Only since 1989 has RNA editing been known to exist in plant mitochon dria. The C-U conversion occurs by desamination, but the mechanisms of other nucleotide exchanges and insertions are still not fully resolved. Many questions are still open. What mechanism is used for RNA editing? Also, the question of the physiological meaning of RNA editing is still unanswered. Is a higher mutation rate of the maternally inherited mt-genome corrected by the editing? From where does the information for the proper nucleotide sequence of the mRNA come? Is this information provided by the nucleus or is it also contained in the mitochondrial genome? It is feasible that the very large mitochondrial genome contains, in addition to the structural genes, sin gle fragments, the transcripts of which are utilized for the correction of the mRNA, as has been observed in the mitochondria from trypanosomes.
Male sterility of plants caused by the mitochondria is an important tool in hybrid breeding
When two selected inbreeding lines are crossed, the resulting F1 hybrids are normally larger, are more robust, and produce higher yields of har vest products than the parent plants. This effect, called hybrid vigor, was observed before the rediscovery of the Mendelian laws of inheritance and was first utilized in 1906 for breeding hybrid maize by George Schull in the laboratory at Cold Spring Harbor in the United States. The success of these studies brought about a revolution in agriculture. Based on the results of Schull’s research, private seed companies bred maize F1 hybrids that pro vided much higher yields than the customary varieties. In 1965, 95% of the maize grown in the Corn Belt of the United States was F1 hybrids. The use of F1 hybrids was, to a large extent, responsible for the increase in maize yields per acre by a factor of 3.5 between 1940 and 1980 in the United States. The hybrid technique also brought dramatic yield increases for rice, thus in China the rice harvest was increased from 1975 to 2000 by a factor of 1.8.
F1 hybrids cannot be further propagated, since according to the Mendelian laws the offspring of the F2 generation is heterogeneous. Most of the second-generation (F2) plants have some homozygosity, resulting in yield depression. Each year, therefore, farmers have to purchase new hybrid seed from the seed companies, whereby the seed companies gained a large economical importance. Hybrid breeding has been put to use for the pro duction of many varieties of cultivated species. Taking maize as an exam ple, the following describes the principles and problems of hybrid breeding.
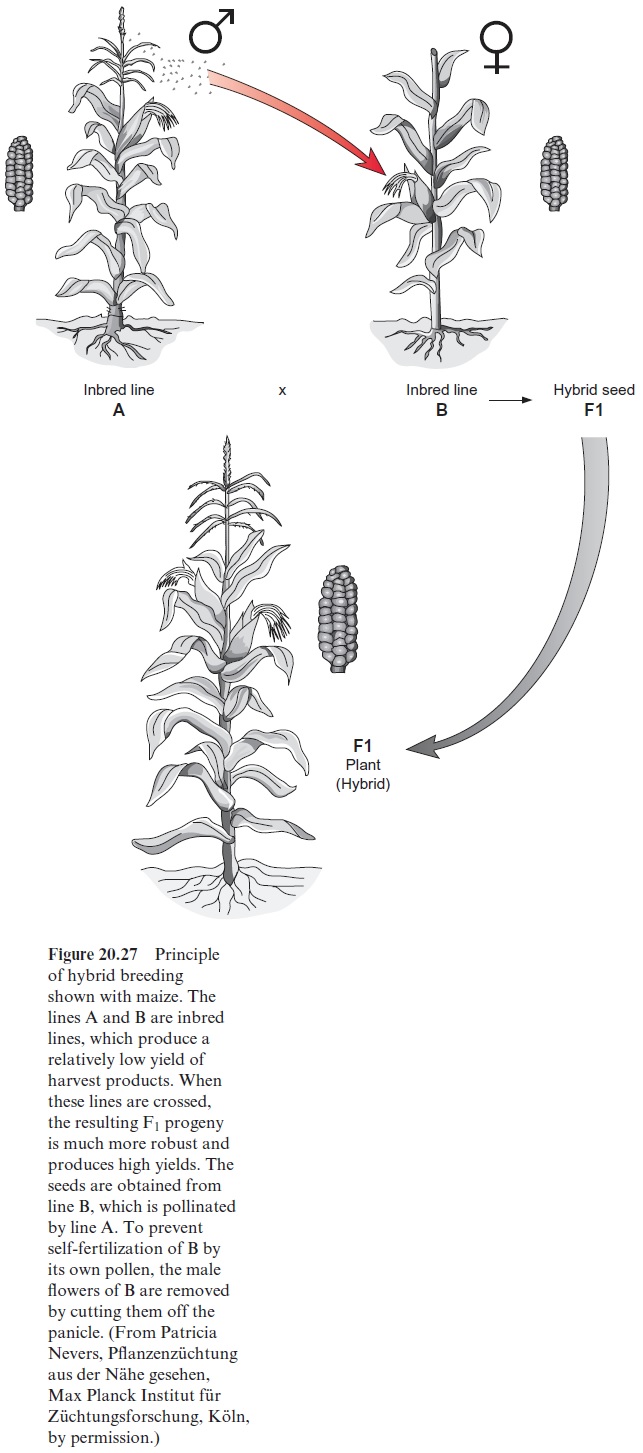
For the production of F1 hybrid seed (Fig. 20.27), the pollen of a pater nal line A is transferred to the pistil of a maternal line B, and only the cobs of B are harvested for seed. These crossings are carried out mostly in the field. Plants of lines A and B are planted in separate, but neighboring rows, so that the pollen is transferred from A to B by the wind. To prevent the pistils of line B from being self- fertilized by the pollen of line B, the plants of line B are emasculated. Since in maize the pollen producing male flowers are separated from the female flowers in a panicle, it is possible to remove only the male flowers by cutting off the panicle. To produce hybrid seeds in this way on a commercial scale, however, requires a great expenditure in manual labor. This method is totally unfeasible on any scale in plants such as rye, where the male and the female parts of the flower are combined.
It was a great step forward in the production of hybrids when maize mutants with sterile pollen were isolated. In these male-sterile plants, the fertility of the pistil was not affected as long as it was fertilized by pollen of other lines. This male sterility is inherited maternally by the genome of the mitochondria. Several male-sterile mutants of maize and other plants are the result of a mutation of mitochondrial genes.
The relationship between the mutation of a mitochondrial gene and the male sterility of a plant has been thoroughly investigated in the maize mutant T (Texas). The mitochondria of this mutant contain a gene desig nated as T-urf13, which encodes a 13 kDa protein. This gene is probably the product of a complex recombination. The 13 kDa protein has no apparent effect on the metabolism of the mitochondria under conditions of vegeta tive growth, and the mutants are of normal phenotypes. Only the formation of pollen is disturbed by this protein, for reasons not known in all details.
It is possible that the tapetum cells of the pollen sac, which are involved in pollen production, have an unusual abundance of mitochondria and appar ently depend very much on mitochondrial metabolism. Therefore, in these cells a mitochondrial defect, which normally does not affect metabolism, might interfere with pollen production.
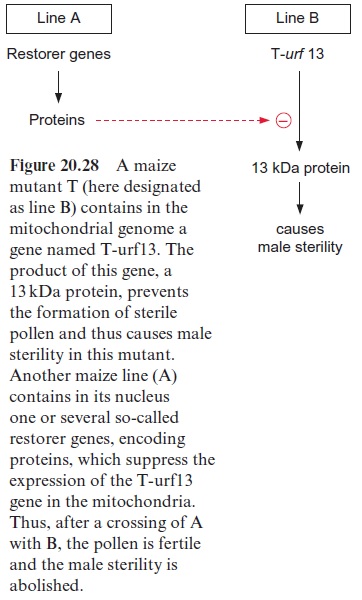

The successful use of these male-sterile mutants for seed production is based on a second discovery: maize lines contain so-called restorer genes in their nucleus. These encodepentatricopeptide repeat proteins (PRR) which are grouped into a protein family that is ubiquitous in plants. Structural prediction indicates that these PPR proteins are present in the organelle and bind RNA, causing the degradation of this RNA. In this way PPR represses the expression of the T-urf13 gene in the mitochondria (Fig. 20.28). The crossing of a paternal plant A containing these restorer genes with a male-sterile maternal plant B results in F1 generation, in which the fertility of the pollen is restored and corn cobs are produced normally.
The crossing of male-sterile T maize lines with lines containing restorer genes enables F1 hybrids to be produced very efficiently. Unfortunately, however, the 13 kDa protein encoded by the T-urf13 makes a maize plant more sensitive to the toxin of the fungus Bipolaris maydis T, the patho gen of the much dreaded fungal disease “southern corn blight,” which destroyed a large part of the American maize crop in 1971. The 13 kDa pro tein reacts with the fungal toxin to form a pore in the inner mitochondrial membrane and thus eliminates mitochondrial ATP production. In order to continue hybrid seed production, it was then necessary to return to the manual removal of the male flowers. Male-sterile lines are now known not only in maize, but also in many other plants, in which the sterility is caused by proteins encoded in the mt-genome, and also other lines that sup press the formation of the inhibiting protein by nuclear encoded proteins. Nowadays these lines are used for the production of fertile F1 hybrid seeds
Presumably the reaction of nuclear encoded proteins on the expression of mitochondrial genes such as T-urf13 is a normal reaction of mitochondrial metabolism in plants and therefore, after the production of corresponding mutants, can be used in many ways to generate male sterility.
Today intensive research is being carried out all over the world to find ways of generating male sterility in plants by genetic engineering. Success can be noted. Using a specific promoter, it is possible to express a ribo nuclease from the bacterium Bacillus amyloliquefaciens exclusively in the tapetum cells of the pollen sac in tobacco and rape seed. This ribonuclease degrades the mRNA formed in tapetum cells, thus preventing the develop ment of pollen. Other parts of the plants are not affected and the plants grow normally. For the generation of a restorer line, the gene of a ribonu clease inhibitor (from the same bacterium) was transferred to the tapetum cells. The great advantage of such a synthetic system is its potential for gen eral application. In this way male sterility can be introduced into species in which this cannot be achieved by manual removal of the stamen, and where male sterility due to mutants is not available. Genetically engineered rape seed hybrids are nowadays grown to a large extent in the United States and Canada. It is to be expected that the generation of male-sterile plants by genetic engineering and the resultant use of hybrid seed might lead to increased harvests of many crops.
Related Topics