Chapter: Biochemistry: Biosynthesis of Nucleic Acids: Replication
The SOS Response in E. coli
The SOS Response in E. coli
When
bacteria are subjected to extreme conditions and a great deal of DNA damage
occurs, the normal repair mechanisms are not up to the task of repairing the
damage. Prolonged exposure to ultraviolet light can do much damage to bacterial
DNA. However, bacteria have one last card to play, which is called,
appropriately, the SOS response. At least 15 proteins are activated as part of
this response, including the mysterious DNA polymerase II. Another important
protein is called recA. It gets its
name from the fact that it is involved in a recombination event. Homologous DNA
can recombine by a variety of mechanisms. Suffice it to say that DNA sequences
exist that can be used to cross one strand over another and replace it. Part
(a) of the figure shows how this might work. If there is a lesion too complex
for the normal repair enzymes to function, a gap is left behind during
replication because DNA polymerases could not synthesize
However, the other replicating strand (shown in blue) should
have the correct complement. RecA and many other proteins recombine this
section of DNA to the lower strand. This leaves the upper strand without a
piece of DNA, but it, too, has its correct complement (shown in red), so DNA
polymerases can replicate it.
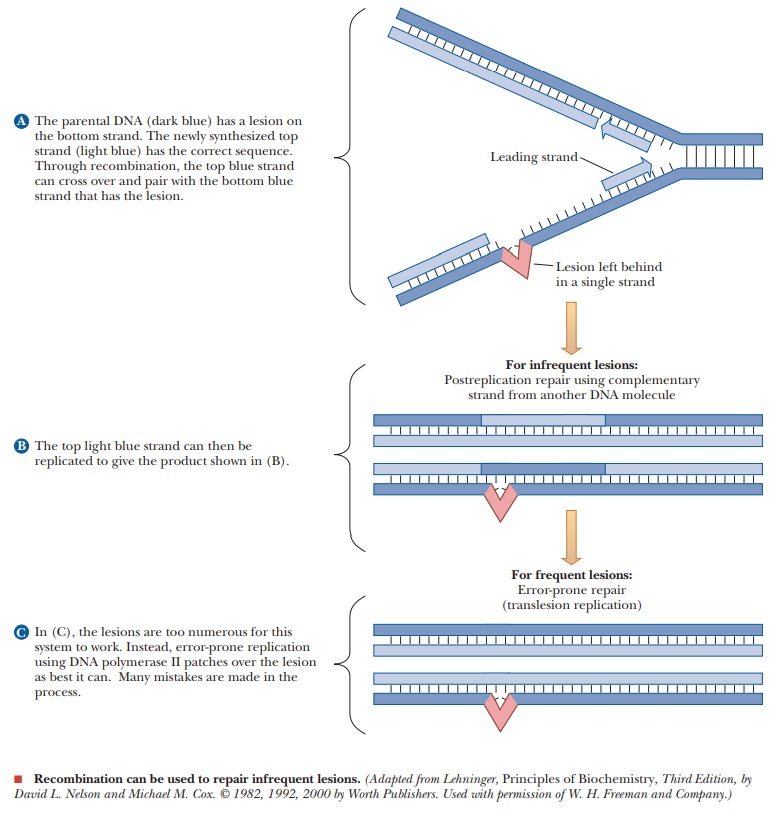
If the damaged strand has too
many lesions, DNA polymerase becomes involved in error-prone repair. In this case, the DNA polymerase continues to
replicate over the damaged area, although it can’t really match bases directly
over the lesions. Thus, it inserts bases without a template, in essence
“guessing.” This goes against the idea of fidelity of replication, but it is
bet-ter than nothing for the damaged cells. Many of the replication attempts produce
mutations that are lethal, and many cells die. However, some may survive, which
is better than the alternative.
Related Topics