Chapter: Genetics and Molecular Biology: Advanced Genetic Engineering
Southern, Northern, and Western Transfers - Genetic Engineering
Southern, Northern, and Western Transfers
Here we will cover in greater detail the topic of
Southern transfers that were mentioned and briefly described. At the same time,
since the concepts are almost the same, we will also mention the so-called
Northern and Western transfers. Southern transfers once were a necessary step
in chromosome mapping, but their use has been superseded by techniques based on
the polymerase chain reaction. To review, DNA fragments can be separated
according to size by electro-phoresis through gels, denatured, transferred to a
nylon or paper mem-brane and immobilized. Then the membrane can be immersed in
a
Figure
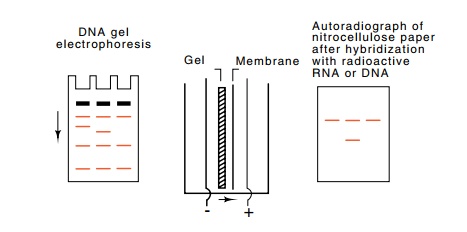
10.5 Southern transfer methodology.
After electrophoretic separationaccording to size, the fragments are denatured
and electrophoretically trans-ferred to a membrane before hybridization with
radioactive probe.
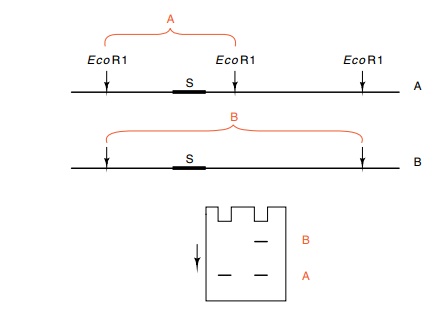
Figure
10.6 Detection of an RFLP by Southern
transfer in which the radioac-tive probe was fragment S. The chromosome A
generates band A and the chromosome B which lacks the first EcoRI cleavage site
on the right generates band B. A heterozygous individual shows both bands in a
Southern transfer probed with the segment S.
buffer containing a labeled oligonucleotide or DNA
fragment and incu-bated under conditions permitting hybridization between
complemen-tary nucleic acid sequences (Fig. 10.5). The labeled fragment will
therefore hybridize to its complementary sequence. The portion of the membrane
carrying the fragment will then become radioactively la-beled, and can be
detected by autoradiography. This part of the process is analogous to plaque
and colony screening described in the previous chapter. This simple technique, named
a Southern transfer for Southern who first devised it, can be used in the
analysis of chromosome structure.
Consider the problem of learning whether the two
nearest EcoRI cleavage sites on
either side of a segment of DNA are at the same location in two nearly
homologous chromosomes. If they are not, the situation is described as a
restriction fragment length polymorphism, RFLP, and the nucleotide differences
producing this polymorphism can be used as a genetic marker. If the restriction
fragment containing the sequence is the same size from both chromosomes, then
the nearest EcoRI cleavage sites are
likely to be in the same locations. To search for RFLPs, a DNA sample
containing the two chromosomes is cut with a restriction en-zyme, separated by
electrophoresis, and “probed” with a radioactively labeled segment of the
region (Fig. 10.6). A difference in the sizes of corresponding fragments
indicates the presence of an RFLP.
Northern transfers are the converse of Southern
transfers in that it is RNA rather than DNA that is separated by
electrophoresis, trans-ferred, and immobilized on membrane to preserve the
original pattern.
Membrane with immobilized RNA can then be used in
hybridization, just like paper with immobilized DNA.
What kinds of questions can be answered with
immobilized RNA? One concerns the in vivo
state of various RNAs. Transient precursors of a mature RNA molecule can easily
be detected because they will be larger than the mature RNA and will be
separated during electrophore-sis. This permits tracking the maturation process
of an RNA molecule. Not only can the changing sizes of the maturing species be
monitored, but fates of specific regions that are removed can also be followed
by probing with appropriate sequences.
Transfer-like technology can also be used to purify
specific RNAs or DNAs. Either single-stranded RNA or single-stranded DNA can be
bound to the paper. Then either RNA or DNA fragments complementary to the
immobilized RNA or DNA can be isolated from a mixture by hybridization followed
by elution. As an application, messenger RNA eluted from such immobilized DNA
can be translated in vitro to provide
a definitive identification of a candidate clone for a specific gene.
Western transfers involve proteins, not nucleic
acids. The principle is the same as for Northern and Southern transfers. A
pattern of proteins that have been separated by electrophoresis is transferred
to paper or a membrane and then specific proteins are visualized. Some DNA- or
RNA-binding proteins can easily be detected after transfer. These proteins
partially renature despite being stuck to the paper. Then the paper with the
immobilized proteins is incubated with the radioactive nucleic acid which binds
to the immobilized protein. After washing the paper to remove unbound
radioactive nucleic acid, autoradiography of the paper reveals the location of
the immobilized protein. More often, the position of a specific protein is
revealed by antibody probing as described in the previous section.
Related Topics