Chapter: Medical Microbiology: An Introduction to Infectious Diseases: Bacterial Genetics
R Plasmids - Bacterial Genetics
R Plasmids
Plasmids that include genes conferring resistance to antimicrobics are of great significance to medicine. They are termed R plasmidsor R factors (resistance factors). The genes responsible for resistance usually code for enzymes that inactivate antimicrobics or reduce the cell’s permeability to them. In contrast, resistance conferred by chromosomal mutation usually involves modification of the target of the antimicrobics (eg, RNA polymerase or the ribosome).
R plasmids occupy center stage in approaches to chemotherapy because of the con-stellation of properties they possess. Those of Gram-negative bacteria can be transmitted across species boundaries and, at lower frequency, even between genera. Many encode re-sistance to several antimicrobics and can thus spread multiple resistance through a diverse microbial population under selective pressure of only one of those agents to which they confer resistance. Nonpathogenic bacteria can serve as a natural reservoir of resistance determinants on plasmids that are available for spread to pathogens.
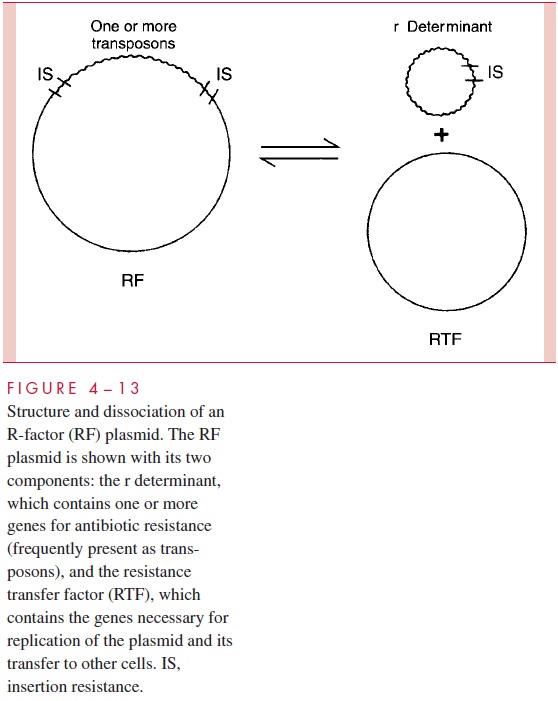
R plasmids evolve rapidly and can easily acquire additional resistance-determining genes from fusion with other plasmids or acquisition of transposons. Many have the capa-bility of amplifying the number of copies of their resistance genes either by gene duplica-tions within each plasmid or by increasing the number of plasmids (copy number) per cell. By these means resistance can be achieved to very high concentrations of the antimi-crobic. One process of gene amplification is based on the ability of some conjugative plasmids to dissociate their components into two plasmids, one (called the resistancetransfer factor) containing genes for replication and for transfer and another (calledthe resistance or r determinant) containing genes for replication and for resistance. Subsequent relaxed replication of the r determinant expands the cell’s capacity to produce the resistance-conferring enzyme (Fig 4 – 13).
Over the past three decades, many of the molecular feats of R plasmids have been explained on the basis of known genetic and evolutionary mechanisms. The discovery of transposable elements (insertion sequences and transposons) and their properties provides an explanation for many of these phenomena. Most plasmids, and all R factors, contain many IS elements and transposons. In fact, virtually all the resistance determinant genes on plasmids are present as transposons. As a result, these genes can be amplified by tan-dem duplications on the plasmid and can hop to other plasmids (or to the bacterial chro-mosome) in the same cell. Combined with the natural properties of many plasmids to transfer themselves by conjugation (even between dissimilar bacterial species), the rapid evolutionary development of multiple drug resistance plasmids and their spread through populations of pathogenic bacteria during the past three decades can be seen as a pre-dictable result of natural selection resulting from the widespread and intensive use of an-timicrobics in human and veterinary medicine .
The properties of transposons can explain the present-day ubiquity and mobility of resistance genes but not their origin. Two facts help point to at least a direction in which to search for an answer. First, R plasmids carrying the genes encoding antimicrobic-inactivating enzymes have been found in bacterial cultures preserved by lyophilization (freeze-drying) since before the era of antimicrobic therapy; an accelerated evolutionary development need not be invoked. Second, the enzymes themselves are remarkably simi-lar to those found in certain bacteria (Streptomyces spp) that produce many clinically useful antimicrobics. Perhaps a long time ago there was a cross-genus transfer of genetic information (by transformation?) that became stabilized on plasmids under the selection pressure of an antimicrobic released into the environment under natural conditions.
Related Topics