Chapter: Medical Microbiology: An Introduction to Infectious Diseases: Bacterial Genetics
Mutation and Repair - Bacterial Genetics
MUTATION AND REPAIR
The spontaneous development of mutations is a major factor in the evolution of bacteria. Mutations occur in nature at a low frequency, on the order of one mutation in every mil-lion cells for any one gene, but the large size of microbial populations ensures the pres-ence of many mutants.
Kinds of Mutations
Mutations are heritable changes in the structure of genes. The normal, usually active, formof a gene is called the wild-type allele; the mutated, usually inactive, form is called the mu-tant allele. There are several kinds of mutations, based on the nature of the change in nu-cleotide sequence of the affected gene(s). Replacements involve the substitution of one base for another. Microdeletions and microinsertions involve the removal and addition, respectively, of a single nucleotide (and its complement in the opposite strand). Insertions involve the addition of many base pairs of nucleotides at a single site. Deletions remove a contiguous segment of many base pairs. Inversions change the direction of a segment of DNA by splicing each strand of the segment into the complementary strand. Duplications produce a redundant segment of DNA, usually adjacent (tandem) to the original segment.
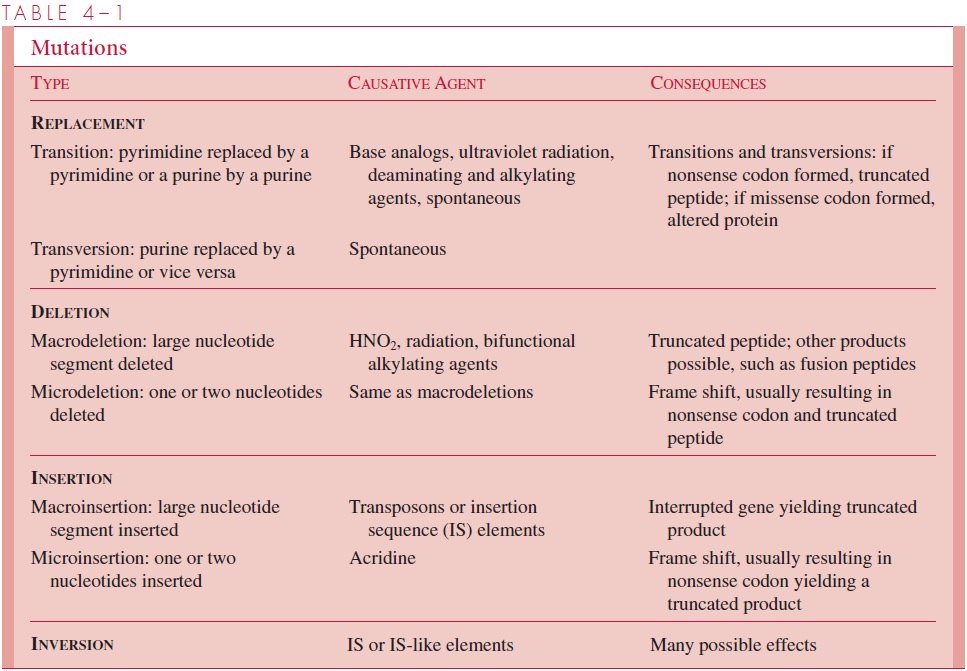
By recalling the nature of genes and how their nucleotide sequence directs the synthe-sis of proteins, one can understand the immediate consequence of each of these biochemi-cal changes. If a replacement mutation in a codon changes the mRNA transcript to a different amino acid, it is called amissense mutation (eg, an AAG [lysine] to a GAG [glutamate]). The resulting protein may be enzymatically inactive or very sensitive to en-vironmental conditions, such as temperature. If the replacement changes a codon specify-ing an amino acid to one specifying none, it is called anonsense mutation (eg, a UAC [tyrosine] to UAA [STOP]), and the truncated product of the mutated gene is called a nonsense fragment.Microdeletions and microinsertions cause frame shift mutations, changes in the reading frame by which the ribosomes translate the mRNA from the mu-tated gene. Frame shifts usually result in polymerization of a stretch of incorrect amino acids until a nonsense codon is encountered, so the product is usually a truncated polypeptide fragment with an incorrect amino acid sequence at its N terminus. Deletion or insertion of a segment of base pairs from a gene shortens or lengthens the protein product if the number of base pairs deleted or inserted is divisible evenly by 3; otherwise it also brings about the consequence of a frame shift. Inversions of a small segment within a gene inactivate it; inverting larger segments may affect chiefly the genes at the points of inversion. Duplications, probably the most common of all mutations, serve an important role in the evolution of genes with new functions. Mutations are summarized in Table 4 – 1.
Many mutations, particularly if they occur near the end of a gene, prevent the expres-sion of all genes downstream (away from the promoter) of the mutated gene. Such polarmutations are thought to exert their effect on neighboring genes by the termination of transcription of downstream genes when translation of the mRNA of the mutated gene is blocked by a nonsense codon.
There is a certain natural frequency of mutations brought about by errors in replication, but various environmental and biological agents can increase the frequency greatly. Differ-ent types of mutations are increased selectively by different agents, as listed in Table 4 – 1.
Mutations may also be classified according to their biological consequences. Some mutations change the susceptibility of a cell to an antimicrobic or other toxic agent; these resistance mutations might, for example, affect the structure of certain cell proteins insuch a way that the agent cannot enter the cell or cannot inactivate its normal target. Some mutations, called auxotrophic mutations, affect the production of a biosynthetic enzyme and result in a nutritional requirement of the mutant cell for the amino acid, nu-cleotide, vitamin, or other biosynthetic product it can no longer make for itself. The wild type from which the mutant was derived is said to be prototrophic for that nutrient. Some mutations affect a gene whose product is essential for growth and cannot be bypassed nutritionally; these are called lethal mutations. If the product of a mutated gene is active in some circumstances but inactive under others (eg, high or low tempera-ture), the mutation is called conditional (meaning conditionally expressed). The most common kind of conditional mutation is one in which the protein product of the mutated gene is inactive at a normally physiologic temperature, but active at a higher or lower temperature; these are called temperature-sensitive mutations.
Reversion and Suppression of Mutations
A reversion, or back mutation, is the conversion of a mutated gene back to its original wild-type allele. True back mutation can occur but at a low frequency, because a very spe-cific and improbable event is required. Much more commonly observed is the conversion of a mutant cell into one that is phenotypically identical to the original wild-type bac-terium for the affected character but still retains the original mutation. These suppressormutations can arise in several ways. Within the mutated codon a second mutation can cre-ate a new codon specifying the original amino acid. Alternatively, secondary mutations in other codons of the mutated gene can lead to a change in amino acid sequence that results in an active product despite the continued presence of the original amino acid error. Suppressing mutations can occur even in genes other than the one that was originally mutated. For example, when two proteins interact to perform a function, the mutant form of one may be active when combined with a mutant form of the other. Another example involves tRNA molecules, the translators of the genetic code, which can themselves be altered by mutation; it is possible for a mutant tRNA to “mistake” a mutant codon and insert the original correct amino acid, a case of two wrongs making a right.
Repair of DNA Damage
Many mutagenic agents directly alter the structure of DNA, and some are ubiquitous components of the environment (heat, sunlight, acid, oxidants, and alkylating agents). It is therefore not surprising to learn that bacteria have evolved multiple biochemical mech-anisms for repairing damaged DNA. In E. coli, for example, more than 30 genes are known to be involved in DNA repair; many of these are members of the SOS response discussed. Collectively these repair systems can remove thymine dimers pro-duced by ultraviolet (UV) irradiation, can remove methyl or ethyl groups placed on gua-nine residues, can excise bases damaged by deamination or ring breakage and replace them with authentic residues, and can recognize and repair DNA depurinated by acid or heat. In large measure these repair systems use the fact that DNA is double stranded. Damage is recognized by the mispairing it causes, and the information on one strand is used to direct the proper repair of the damaged strand. Also, a proofreading process oper-ates during DNA replication to detect any mismatch between each newly polymerized base and its mate in the template strand. Mismatches are excised to permit repolymeriza-tion with the properly matched nucleotide. Failures of this proofreading process can be detected and handled by an excision and resynthesis system similar to those that recog-nize and repair chemically damaged DNA.
One system bypasses DNA damaged by UV irradiation when repair has failed. It directs replication to proceed across a region badly damaged by the formation of thymine dimers. This error-prone replication is responsible for the mutations induced by UV light.
Related Topics