Chapter: Biotechnology: Recombinant DNA Technology
Polymerase Chain Reaction (PCR)
Polymerase Chain Reaction (PCR)
The polymerase chain reaction or PCR as it is commonly known, was invented by Kerry Mullis in 1985. It results in the selective amplification of a specific region of a DNA molecule and so can also be used to generate a DNA fragment for cloning. The basic principle underlying this technique is that when a double-stranded DNA molecule is heated to a high temperature, the two DNA strands separate giving rise to single stranded molecules which can be made to hybridise with small oligonucleotide primers (single-stranded) by bringing down the temperature. If to this an enzyme called DNA polymerase and nucleotide triphosphates are added, much like what happens during replication, i.e primer extension occurs. This procedure is repeated several times, see Fig. 8 which ultimately results in amplification of the DNA stretch between the two primers (one on each strand of the DNA). The basic requirements of a PCR reaction are:
1. DNA template to be amplified.
2. Primers which are oligonucleotides, usually 10-18 nucleotides long that hybridise to the target DNA region one to each strand of the DNA. Two primers of such a sequence are required so that they can hybridise as indicated in the Fig. 8.
3. DNA polymerase which is stable at temperatures above 80oC . Taq polymerase which has been isolated from a thermostable bacterial species is used.
4. Deoxynucleotide triphosphates and buffer.
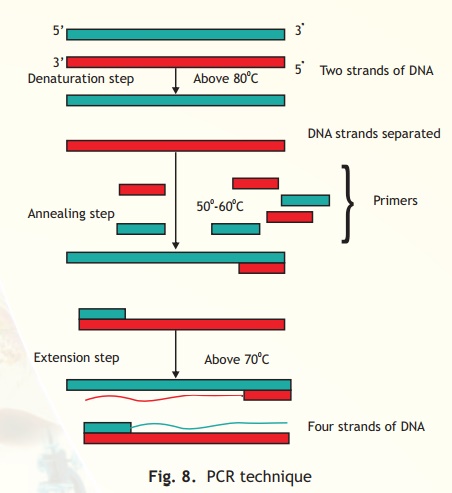
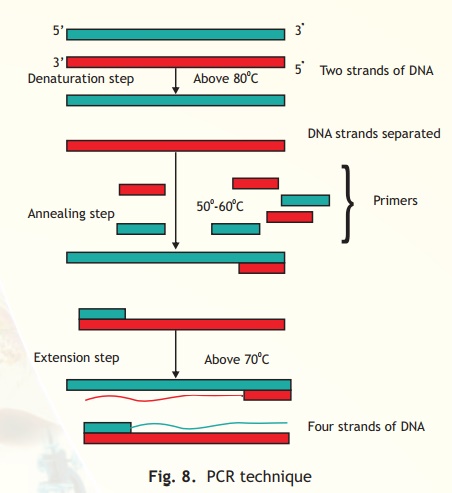
Fig. 8. PCR technique
A single PCR amplification cycle involves three basic steps of denaturation, annealing and extension (Fig. 8). In the denaturing step, the target DNA is heated to a high temperature, above 80oC which results in DNA strand separation. Each single strand then anneals with a primer at a lower temperature between 50-60oC in such a way that extension can occur from it in a 5' - 3' direction (a requirement of the DNA polymerase). The final step is extension, wherein the enzyme Taq polymerase extends each primer using dNTPs and the DNA strand as template. The temperature for extension is around 70oC. The procedure is repeated and each set of steps is considered as one cycle (i.e denaturation, annealing and extension). At the end of one cycle two DNA molecules have become four and this geometric progression occurs with each cycle. Hence it can be computed that at the end of n cycles the number of DNA molecules is 2n which is enormously amplified target DNA.
The invention of the PCR technique has revolutionised every aspect of modern biology. To detect pathogens, microbiologists in the past used techniques based on culturing and detecting antibodies against enzymes or proteins specific to the pathogen. Apart from taking time many of these procedures were not specific. PCR based diagnosis is faster, safer and more specific because it does not use live pathogens; instead DNA from the infected tissue is isolated and the PCR technique is carried out using primers having specific complementary sequences to the pathogen DNA. PCR is also a valuable tool in forensic science as large amounts of DNA can be amplified from the small amounts present at the crime site, for DNA fingerprinting analysis. In recent years, PCR has also found use in detecting specific microorganisms from environmental samples of soil, sediments and water. It is interesting that archaeologists are using combinations of PCR and fingerprinting analysis to relate and establish ancient Egyptian dynasties from samples obtained from mummies.
Related Topics